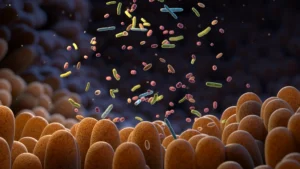
The intricate ecosystem residing within the human gastrointestinal tract, collectively known as the gut microbiome, represents a cornerstone of human physiological well-being. This vast, dynamic consortium of microorganisms, often numbering in the trillions, performs indispensable functions ranging from nutrient metabolism and vitamin synthesis to immune system modulation and protection against pathogens. At the heart of this complex biological partnership lies a continuous, sophisticated dialogue—a chemical exchange occurring not only among the microbial inhabitants themselves but also between these microbes and their human host. For this delicate balance and mutualistic interaction to persist, the individual bacterial constituents must possess an acute awareness of their immediate surroundings, adeptly detecting a multitude of chemical signals and available nutrients. Despite the acknowledged critical importance of these sensory capabilities, the precise array of chemical cues that bacterial receptors can recognize, particularly within the context of beneficial gut species, has remained a significant area of scientific inquiry.
A fundamental question that has long puzzled microbiologists centers on identifying the specific chemical information that holds the most significance for the health-promoting bacteria inhabiting the human gut. Historically, the field of microbiology has predominantly channeled its research efforts into understanding pathogenic organisms—those bacteria responsible for causing disease. This focus, driven by urgent public health imperatives, has yielded profound insights into bacterial virulence mechanisms and strategies for combating infections. Consequently, a disproportionately smaller volume of research has been dedicated to elucidating the sensory mechanisms and ecological roles of commensal microbes—the non-pathogenic, often beneficial microorganisms that naturally colonize the human body without causing harm, and frequently providing tangible benefits. This research disparity has created a substantial knowledge gap, leaving scientists with limited understanding of how these helpful bacterial residents genuinely perceive and react to the diverse chemical landscape within their host’s digestive system.
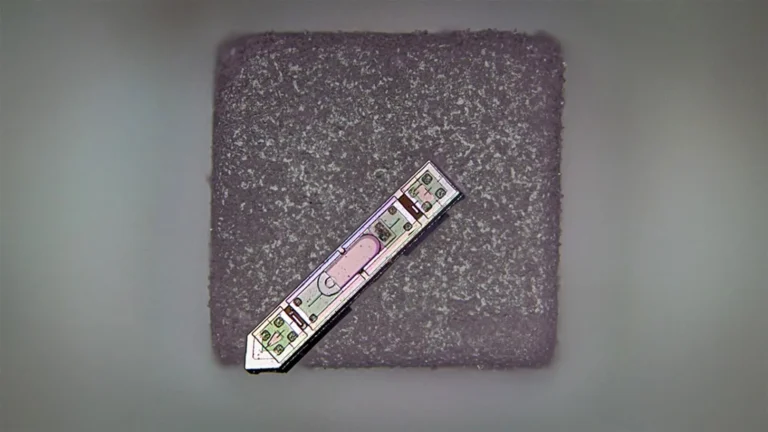
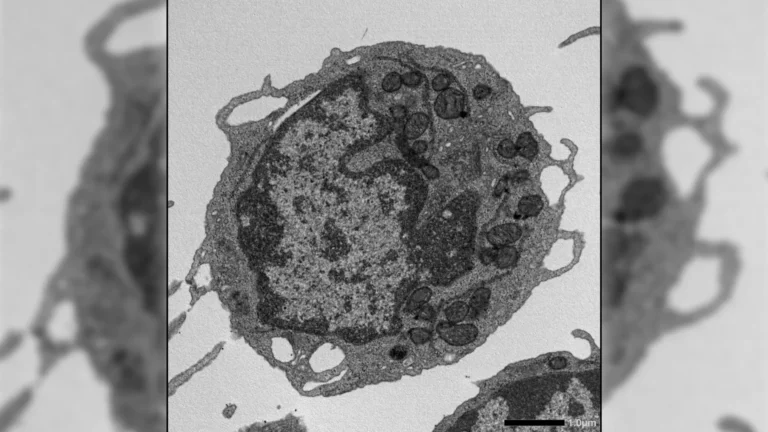
Addressing this critical void, a pioneering international research collaboration embarked on an ambitious quest to unravel the sensory preferences of beneficial gut bacteria. This distinguished team, comprising scientists from the Max Planck Institute for Terrestrial Microbiology, the University of Ohio, and the Philipps-University Marburg, focused its investigation on the genus Clostridia. These bacteria are recognized as abundant and highly motile inhabitants of the human gut, playing a crucial role in maintaining intestinal health, including the production of beneficial short-chain fatty acids (SCFAs) like butyrate, which nourish gut lining cells and modulate immune responses. The selection of motile bacteria like Clostridia was strategic, as their ability to move actively implies a sophisticated sensory system enabling them to navigate towards favorable conditions or away from detrimental ones, making them ideal subjects for studying chemical detection.
Through a meticulous and systematic screening process, the researchers made a groundbreaking discovery: receptors originating from the human gut microbiome possess the capacity to recognize an astonishingly diverse spectrum of metabolic compounds. These crucial substances encompass breakdown products derived from major macronutrients such as carbohydrates, fats, and proteins, alongside components like DNA fragments and various amines. This extensive recognition capability underscored the bacteria’s sophisticated ability to survey their environment for essential resources. Furthermore, the comprehensive analysis revealed distinct patterns in these sensory preferences. Different classes of bacterial sensors exhibited clear and specific affinities for particular categories of chemicals, indicating that the gut bacteria are not merely reacting indiscriminately to their environment. Instead, they are finely tuned, selectively responding to specific metabolic signals that are pertinent to their survival, growth, and overall ecological function within the host. This selective tuning highlights an underlying biological intelligence guiding bacterial behavior.
Among the myriad chemical ligands identified that bind to the sensory receptors controlling bacterial movement, lactic acid (lactate) and formic acid (formate) emerged with remarkable frequency as primary stimuli. This compelling finding suggests that these two compounds may serve as exceptionally important nutrient sources and signaling molecules for a broad range of gut bacteria. The research, which ingeniously combined rigorous laboratory experimentation with advanced bioinformatic analysis, demonstrated that bacterial motility is predominantly driven by the active pursuit of food. These receptors act as molecular antennae, guiding the bacteria towards patches of higher nutrient concentration, thereby optimizing their access to vital resources. The prevalence of lactate and formate as key attractants provides critical insights into the energetic landscape of the gut, pointing to their significant role as metabolic intermediates within the complex food web of the microbiome.
The prominence of lactate and formate also illuminates the vital ecological concept of "cross-feeding," a cornerstone of a stable and resilient gut ecosystem. In this cooperative process, certain bacterial species generate metabolites, such as lactate and formate, as byproducts of their own metabolic activities. These metabolites, rather than being waste products, are then utilized as essential food sources by other distinct bacterial species within the community. This metabolic interdependence fosters a remarkable level of efficiency and resource partitioning, minimizing competition and maximizing the overall productivity of the microbial community. As Dr. Wenhao Xu, a postdoctoral researcher in Victor Sourjik’s group and the study’s first author, explained, "These domains appear to be important for interactions between bacteria in the gut and could play a key role in the healthy human microbiome." This intricate web of metabolic exchange is fundamental to the stability of the gut environment, ensuring that the collective community thrives and, in turn, contributes to the health of the human host. Disruptions to this delicate balance can have profound implications for overall health, underscoring the importance of understanding these foundational interactions.
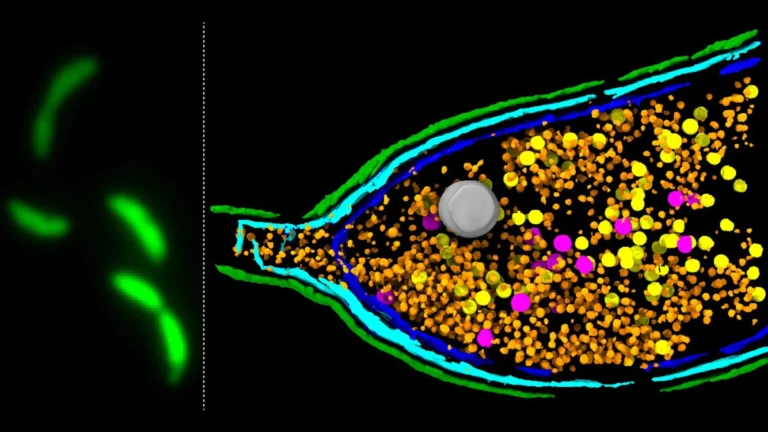
Beyond identifying preferred nutrients, the research team made another significant breakthrough: the discovery of several previously unknown groups of sensory domains. These newly characterized sensors exhibit specific responsiveness to lactate, dicarboxylic acids, uracil (a crucial building block of RNA), and short-chain fatty acids (SCFAs). The identification of sensors for SCFAs, such as acetate, butyrate, and propionate, is particularly noteworthy, given their established critical roles in host energy metabolism, immune regulation, and gut barrier integrity. Further advancing their understanding, the scientists successfully determined the precise crystal structure of a newly identified "dual sensor" capable of responding to both uracil and acetate simultaneously. This detailed molecular-level insight revealed how these distinct molecules bind to specific sites on the sensor, elucidating the physical mechanisms of recognition. The existence of such a dual-function sensor highlights evolutionary efficiency, allowing a single molecular apparatus to monitor multiple, potentially related, environmental cues, and this sensor belongs to a vast family of sensory domains, suggesting a widespread and adaptable system of chemical detection in bacteria.
The investigation also provided compelling evidence for the remarkable evolutionary flexibility of bacterial sensory systems. By meticulously examining the phylogenetic relationships among uracil sensors and other related sensory domains, the team uncovered that the specificity of a ligand—the molecule a sensor binds to—can undergo relatively facile changes over evolutionary time. This inherent adaptability is a critical feature that enables bacteria to rapidly fine-tune their sensing capabilities in response to fluctuating environmental conditions, such as dietary shifts, antibiotic exposure, or changes in host physiology. This capacity for evolutionary adaptation ensures the long-term survival and resilience of bacterial populations within dynamic ecological niches like the human gut.
As Victor Sourjik, the lead researcher, aptly summarized, "Our research project has significantly expanded the understanding of sensory abilities of beneficial gut bacteria. To our knowledge, this is the first systematic analysis of the sensory preferences of non-model bacteria that colonize a specific ecological niche." He further emphasized the broader implications, stating, "Looking ahead, our approach can be similarly applied to systematically investigate sensory preferences in other microbial ecosystems." The profound insights gleaned from this study not only deepen our fundamental understanding of how beneficial gut bacteria perceive and interact with their environment but also pave the way for future research into manipulating these sensory pathways. Such advancements could potentially lead to innovative strategies for promoting a healthy microbiome, preventing disease, and developing next-generation probiotics or therapeutic interventions aimed at precisely modulating the gut’s intricate chemical dialogue for improved human health outcomes.