Scientists have elucidated a fascinating and previously misunderstood mechanism by which bacteria actively dismantle themselves to facilitate the dissemination of genetic material, including crucial genes that confer resistance to antibiotics, a phenomenon posing a significant and escalating threat to global public health. Researchers at the John Innes Centre, in collaboration with esteemed institutions including the University of York and the Rowland Institute at Harvard, have delved into the intricate workings of enigmatic entities known as gene transfer agents (GTAs). These microscopic particles, superficially resembling bacteriophages – viruses that infect bacteria – have undergone a remarkable evolutionary transformation. Once foreign invaders, they have been domesticated by bacteria, effectively repurposed and brought under the host cell’s direct command.
These sophisticated GTA particles function as highly specialized biological couriers, meticulously collecting segments of DNA from within a parent bacterial cell. Subsequently, they embark on a journey to deliver these genetic payloads to neighboring bacteria, a process critically important for bacterial evolution and adaptation. This form of genetic exchange, termed horizontal gene transfer, enables bacteria to rapidly acquire and propagate advantageous traits. Among the most significant of these traits are genes that endow bacteria with the ability to withstand the effects of antimicrobial drugs, thereby rendering infections more difficult to treat.
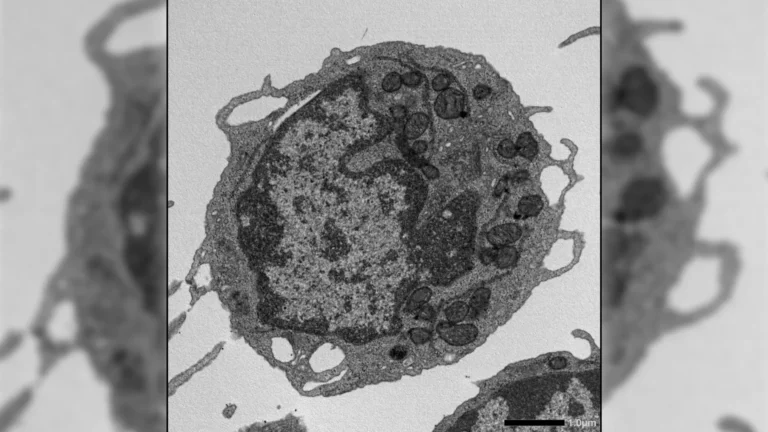
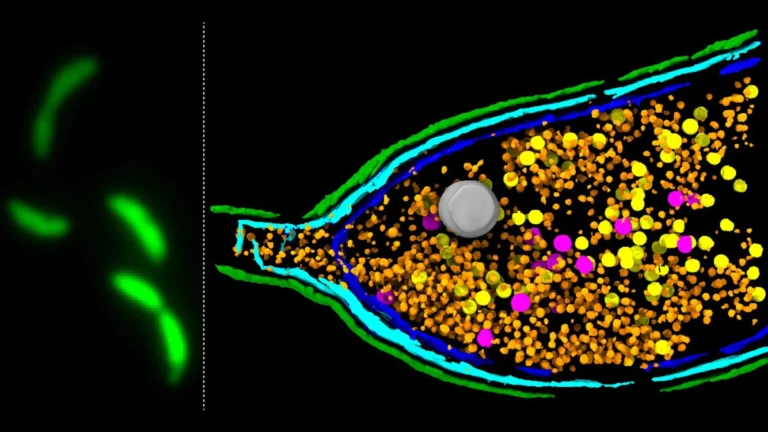
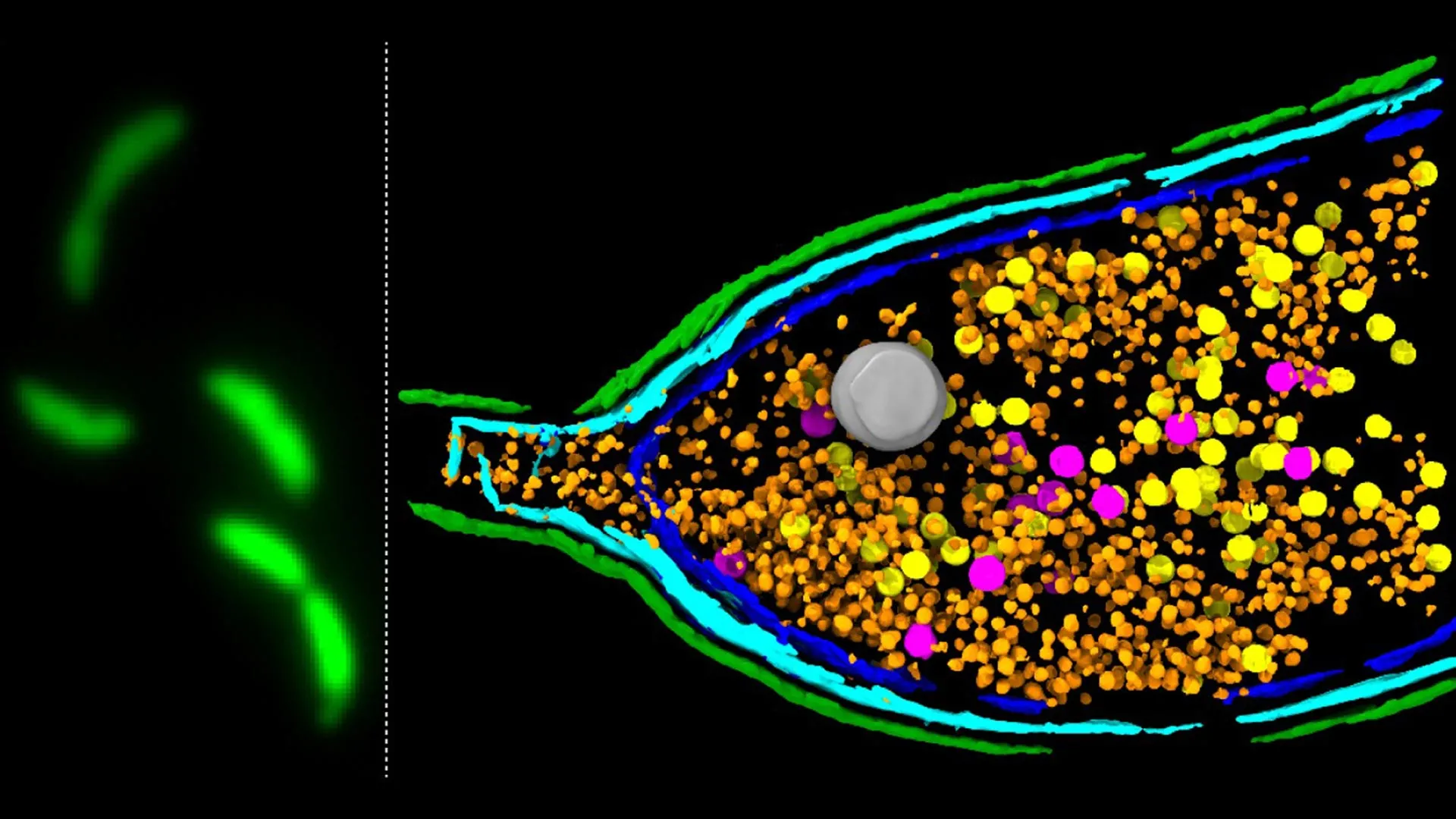
A pivotal and until now, poorly understood, stage in the life cycle of GTAs is the process of host cell lysis. This refers to the controlled rupture or bursting of the bacterial cell membrane, a necessary event that liberates the assembled GTA particles, allowing them to embark on their mission of genetic transfer. Without this fundamental step, the efficient spread of genetic information would be severely hampered.
The groundbreaking research, detailed in the prestigious journal Nature Microbiology, employed a sophisticated deep sequencing-based screening methodology. This advanced technique allowed the scientists to systematically identify the specific genes responsible for driving GTA activity within the well-studied model bacterium Caulobacter crescentus. Their investigation zeroed in on a tripartite genetic locus, designated LypABC, which orchestrates the production of key bacterial proteins.
Crucially, when the genes comprising the LypABC system were experimentally inactivated or removed, the bacterial cells lost their capacity to undergo lysis and consequently, could no longer release GTA particles. Conversely, when the activity of the LypABC system was artificially amplified, a substantial proportion of the bacterial population exhibited premature and widespread cell death through lysis. These compelling experimental outcomes firmly established LypABC as a central regulatory nexus governing the process of cell disintegration for GTA release.
Perhaps one of the most unexpected revelations from this study is the striking molecular resemblance of the LypABC system to established bacterial defense mechanisms, specifically anti-phage immune systems. The protein components encoded by LypABC share significant structural and functional similarities with proteins typically employed by bacteria to ward off viral infections. However, in this novel context, this ancient immune machinery appears to have been ingeniously co-opted and repurposed by the bacteria themselves. Instead of serving a defensive role, it now actively facilitates the release of GTA particles, thereby promoting the very gene transfer that can lead to the spread of beneficial traits, including antibiotic resistance. This finding underscores the remarkable evolutionary flexibility of bacterial biological systems, demonstrating their capacity to adapt and reassign existing molecular machinery for entirely new purposes.
Furthermore, the research team identified an additional regulatory protein that plays a vital role in maintaining stringent control over GTA activity and the associated lysis process. This meticulous regulation is not merely an incidental aspect but a critical requirement for bacterial survival. The improper or uncontrolled activation of the LypABC system can prove exceptionally toxic to the bacterial cell, leading to its demise without the intended benefit of genetic dissemination. This regulatory oversight ensures that the potentially lethal process of cell lysis is tightly managed, occurring only when and where it serves the bacteria’s evolutionary interests.
By illuminating the adaptability and plasticity of bacterial genetic and cellular mechanisms, this study provides profound new insights into the complex pathways by which genes migrate between bacterial populations. This understanding is of paramount importance for comprehending the escalating global challenge of antimicrobial resistance (AMR). The ability of bacteria to rapidly acquire resistance genes through horizontal gene transfer significantly complicates treatment strategies and contributes to the emergence of multidrug-resistant pathogens.
Dr. Emma Banks, the lead author of the study and a Royal Commission for the Exhibition of 1851 Research Fellow, articulated the significance of these findings, stating, "What’s particularly interesting is that LypABC looks like an immune system, yet bacteria are using it to release GTA particles. It suggests that immune systems can be repurposed to help bacteria share DNA with each other – a process that can contribute to the spread of antibiotic resistance." This observation highlights a fascinating evolutionary paradox where a system designed for defense is turned into an instrument for propagation.
The next critical phase of this research endeavor will focus on unraveling the intricate molecular triggers and signaling pathways that initiate the activation of the LypABC control hub. Scientists aim to further elucidate the precise mechanisms by which this system governs the precise timing and controlled rupture of bacterial cells, thereby orchestrating the efficient release of GTA particles. Understanding these fine-tuned regulatory processes could potentially open new avenues for therapeutic intervention, perhaps by disrupting this critical step in the spread of resistance genes.
The implications of this discovery extend beyond the immediate understanding of GTA function. It paints a clearer picture of the dynamic and often surprising evolutionary strategies employed by bacteria. These "enemies-turned-allies," as they can be viewed in the context of gene transfer, demonstrate a remarkable capacity for innovation, leveraging existing cellular machinery to adapt and thrive in diverse environments, including those fraught with antibiotic challenges. The ongoing investigation into these bacterial self-destruction and gene-sharing mechanisms offers crucial clues in the global battle against antibiotic resistance, a fight that demands a deep and comprehensive understanding of bacterial biology at its most fundamental level.