The tenacity of microbial life is a well-documented phenomenon, with bacteria demonstrating an extraordinary capacity to flourish in environments that would be considered inhospitable to most other organisms. These hardy survivors inhabit realms ranging from the searing heat of hydrothermal vents to the profound chill of sub-zero glacial formations. Subterranean ice caves, characterized by their perpetually frozen conditions, represent one such extreme niche, harboring a diverse consortium of microorganisms whose genetic makeup and adaptive strategies are only now coming under scientific scrutiny. These frigid, ancient repositories are believed to contain a vast and largely uncatalogued reservoir of genetic material, offering a unique window into the evolutionary history of life.
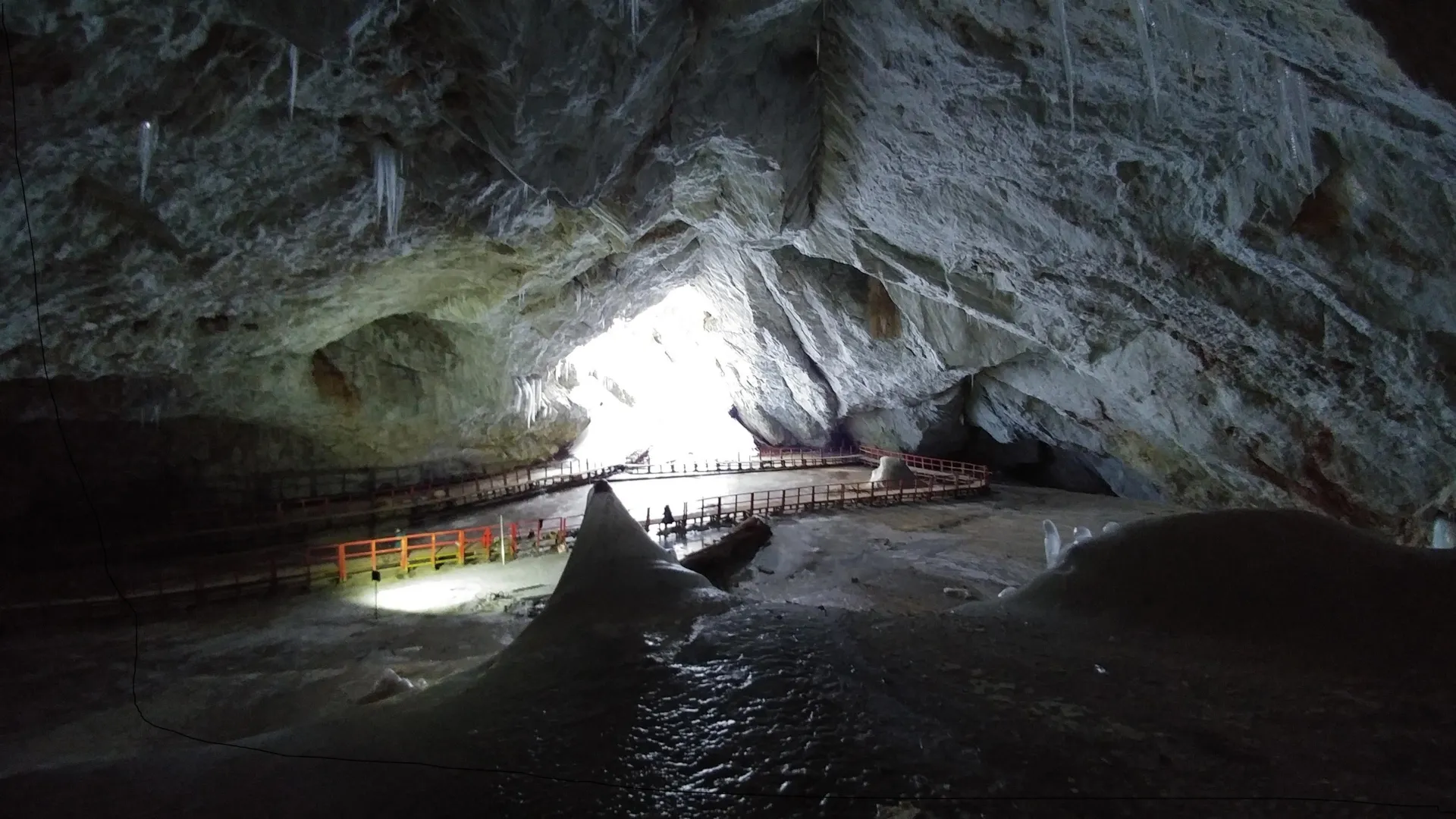
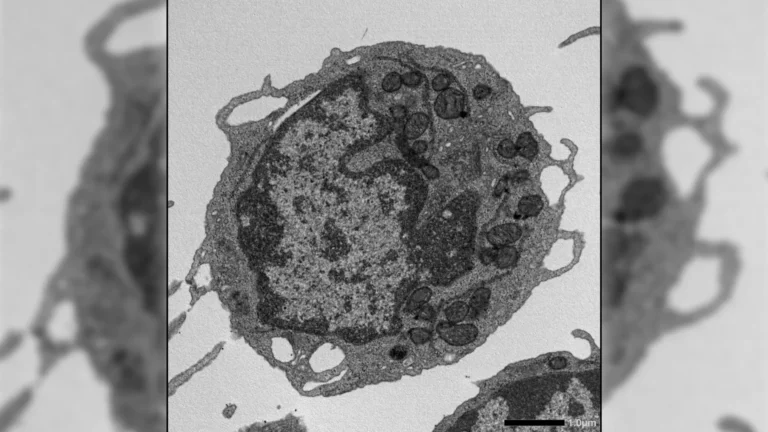
Within the subterranean depths of Romania’s Scarisoara Ice Cave, a team of researchers has meticulously investigated a bacterial strain that has remained entombed for approximately five millennia within a preserved ice layer. Through a comprehensive analysis of its resistance mechanisms against modern pharmacological agents, this ancient microbe has revealed critical information regarding the natural genesis and dissemination of antibiotic resistance. The groundbreaking discoveries stemming from this research have been formally documented and published in the esteemed scientific journal Frontiers in Microbiology.
Dr. Cristina Purcarea, a senior scientist at the Institute of Biology Bucharest of the Romanian Academy, elaborated on the significance of the findings, stating that the bacterial isolate, identified as Psychrobacter SC65A.3, exhibits a remarkable resilience to a multitude of contemporary antibiotics, despite its ancient lineage. Further investigation revealed that this microorganism possesses over 100 genes associated with resistance mechanisms. Intriguingly, this ancient bacterium also demonstrates a capacity to impede the proliferation of several highly problematic, antibiotic-resistant pathogenic bacteria, often referred to as "superbugs." Moreover, the strain exhibits significant enzymatic activities, suggesting considerable potential for applications in various biotechnological fields.
The study of ancient microorganisms, particularly those retrieved from millennia-old ice deposits such as those found in Scarisoara Ice Cave, provides an invaluable opportunity to comprehend the evolutionary trajectory of antibiotic resistance. This resistance emerged naturally within environmental contexts, predating the advent of human-developed antibiotics by a considerable margin. Dr. Purcarea emphasized that by examining such microbes, scientists can gain a deeper understanding of the historical development and natural spread of these resistance traits.
The isolation of the organism involved a precise operation: the research team extracted an ice core extending 25 meters in depth from a section of the cave known as the Great Hall. This core served as a frozen chronicle, potentially encapsulating a record of environmental conditions spanning up to 13,000 years. Rigorous protocols were implemented to preclude any possibility of contamination during the retrieval process. Ice samples were hermetically sealed in sterile containment bags and transported to the laboratory under continuously frozen conditions. Upon arrival, scientists meticulously isolated various bacterial strains. Subsequently, their genomes were subjected to comprehensive sequencing, a process designed to identify genes responsible for survival in extreme cold, as well as those intrinsically linked to antimicrobial resistance and activity.
Following the isolation and genomic analysis, the researchers subjected the Psychrobacter SC65A.3 strain to an extensive battery of tests, exposing it to 28 different antibiotics categorized across 10 distinct classes. These particular antibiotics are frequently prescribed or reserved for the treatment of severe bacterial infections in clinical settings. The research team leveraged existing knowledge of known resistance genes and mutations associated with some of these drugs, enabling them to draw direct comparisons between predicted resistance mechanisms and the observed laboratory outcomes. Dr. Purcarea highlighted that the 10 antibiotics to which the bacterium demonstrated resistance are widely employed in both oral and injectable therapeutic regimens used to manage a spectrum of serious bacterial infections encountered in clinical practice. Notable among these medications were rifampicin, vancomycin, and ciprofloxacin, drugs critical in the treatment of conditions such as tuberculosis, Clostridioides difficile colitis, and urinary tract infections (UTIs).
The Psychrobacter SC65A.3 strain stands out as the inaugural Psychrobacter species identified to exhibit resistance to specific antibiotics, including trimethoprim, clindamycin, and metronidazole. These particular drugs are commonly utilized for the treatment of UTIs and infections affecting vital organs and systems such as the lungs, skin, bloodstream, and reproductive tract. The resistance profile observed in this strain strongly suggests that bacteria adapted to cold environments can function as significant reservoirs for resistance genes. These genes, essentially segments of DNA, confer upon bacteria the ability to survive exposure to antibiotic compounds.
The implications of this discovery present a dual nature, encompassing both potential risks and significant opportunities for scientific advancement. Dr. Purcarea cautioned that the potential release of these ancient microbes through melting ice could facilitate the transfer of their resistance genes to contemporary bacterial populations, thereby exacerbating the escalating global crisis of antibiotic resistance. Conversely, these ancient organisms produce unique enzymes and antimicrobial compounds that hold considerable promise as inspiration for the development of novel antibiotics, industrial enzymes, and other innovative biotechnological applications.
A detailed genetic analysis of Psychrobacter SC65A.3 unveiled a substantial number of genes, nearly 600 in total, whose functions remain as yet undetermined. This finding underscores the immense, largely untapped potential of these ancient microbes as a source for uncovering novel biological processes and mechanisms. Furthermore, the research team identified 11 genes within the bacterial genome that possess the demonstrated capacity to either directly eliminate or inhibit the growth of bacteria, fungi, and even viruses.
As the phenomenon of antibiotic resistance continues its relentless rise on a global scale, insights derived from the study of ancient microbial communities are becoming increasingly indispensable. The examination of genomes meticulously preserved within ice matrices offers scientists a critical tool for tracing the historical emergence and propagation of resistance mechanisms, long before the advent of modern medical interventions. Dr. Purcarea concluded by emphasizing the profound importance of these ancient bacteria to both scientific understanding and medical progress, while simultaneously stressing the absolute necessity for stringent handling protocols and robust safety measures within laboratory settings to effectively mitigate the risk of any uncontrolled dissemination.