A groundbreaking computational investigation by public health researchers at Columbia University’s Mailman School of Public Health has meticulously reconstructed the swift and pervasive spread of two significant respiratory pandemics – the 2009 H1N1 influenza strain and the 2020 SARS-CoV-2 virus – across the diverse urban landscape of the United States. This pioneering study, published in the esteemed journal Proceedings of the National Academy of Sciences, stands as the inaugural direct comparative analysis of how these distinct viral agents infiltrated and propagated through American metropolitan centers. The findings illuminate the astonishing rapidity with which airborne pathogens can achieve widespread distribution and underscore the profound challenges inherent in containing such outbreaks during their nascent stages.
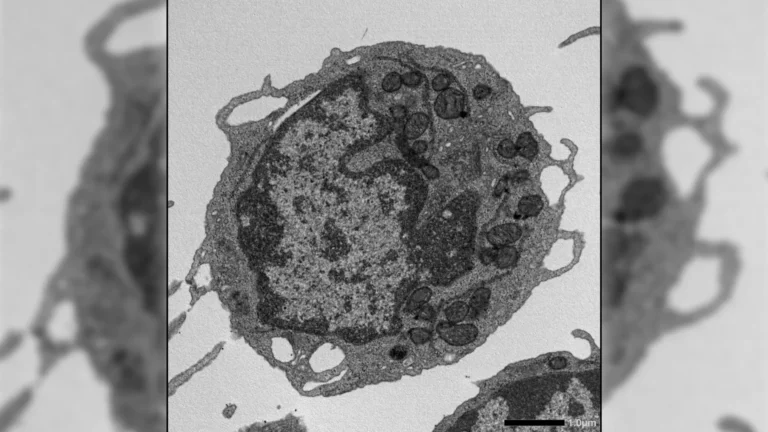
The societal and health ramifications of both pandemics were undeniably substantial. The 2009 H1N1 influenza pandemic resulted in an estimated 274,304 hospitalizations and claimed the lives of 12,469 individuals within the U.S. The subsequent COVID-19 pandemic, however, proved to be a far more catastrophic event, with official reports indicating over 1.2 million confirmed fatalities to date. These figures represent only a fraction of the total burden, as they typically exclude indirect deaths and the long-term health sequelae experienced by survivors. The sheer scale of these events necessitated a deeper understanding of their transmission dynamics, prompting the Columbia University team to delve into the intricate mechanisms of their geographical expansion.
At the heart of this research lies a sophisticated modeling approach designed to unravel the complex interplay of factors driving pandemic dissemination. The scientists integrated detailed epidemiological data pertaining to the transmissibility characteristics of both the H1N1 virus and the SARS-CoV-2 virus with advanced computational simulations. These models meticulously accounted for a multifaceted network of human movement, encompassing both long-distance air travel and the daily ebb and flow of local commuting patterns, while also acknowledging the significant impact of super-spreading events, which can disproportionately amplify transmission. The analytical focus encompassed over three hundred distinct metropolitan areas, providing a comprehensive national perspective.
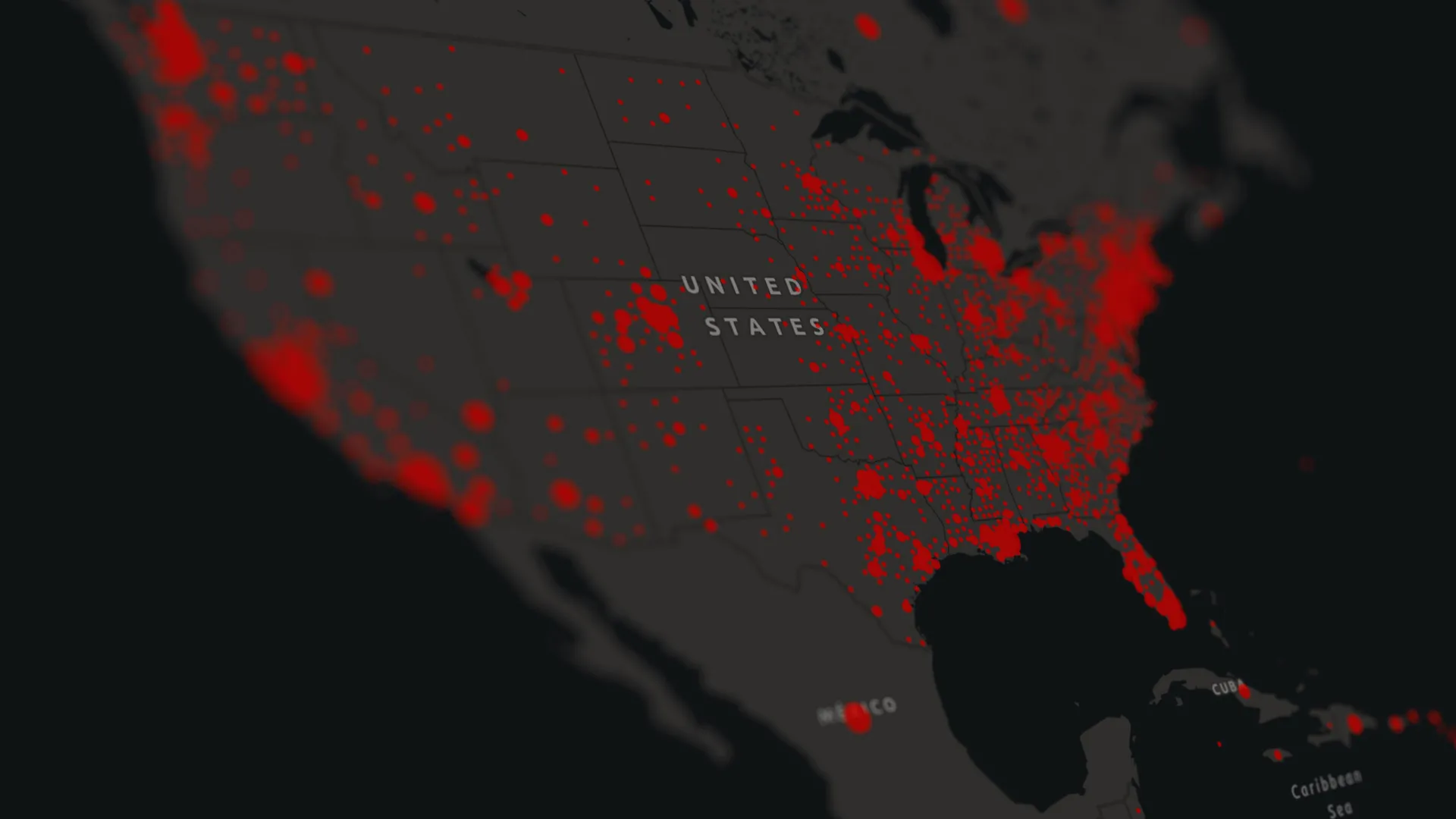
A stark and recurring theme emerging from the simulations is the alarming speed at which these pathogens establish a significant foothold within population centers. The models revealed that by the time early detection mechanisms or public health interventions were effectively implemented, both pandemics had already achieved a substantial degree of circulation across the majority of studied metropolitan regions. This phenomenon highlights a critical window of vulnerability, where the virus can spread extensively before its presence is even widely recognized. While the specific pathways of inter-city transmission differed between H1N1 and COVID-19, a consistent pattern of reliance on major transportation and population hubs, such as New York City and Atlanta, was observed as crucial conduits for viral propagation. The research definitively established that air travel served as a far more potent accelerant for rapid, long-distance spread than the more localized effects of daily commuting. Compounding these predictable patterns of movement was the inherent unpredictability of transmission events themselves, including the influence of superspreaders, which introduced a significant degree of uncertainty, making real-time forecasting of emergent outbreaks exceedingly difficult.
Dr. Sen Pei, an assistant professor of environmental health sciences at the Columbia Mailman School and the study’s senior author, emphasized the critical lessons learned from these events. "The swift and inherently uncertain propagation of both the 2009 H1N1 influenza and the 2020 COVID-19 pandemics starkly illustrates the formidable challenges associated with timely detection and effective control measures," Dr. Pei stated. He further posited that "expanding the reach of wastewater surveillance systems, when coupled with robust and effectively implemented infection control strategies, holds considerable promise for mitigating the initial phases of future pandemic spread."
The concept of wastewater surveillance as an early warning system for infectious disease outbreaks has garnered increasing attention in recent years, and this latest research provides further compelling evidence of its utility. By analyzing sewage samples, public health officials can detect the presence of viral genetic material, offering a population-level indicator of infection trends that often precedes widespread clinical case reporting. This study reinforces the notion that a more extensive and integrated network of wastewater monitoring could serve as an invaluable tool in enhancing overall pandemic preparedness and, crucially, in slowing the initial, most vulnerable stages of transmission.
Beyond the specific dynamics of H1N1 and COVID-19, the researchers have developed a versatile analytical framework with broader applicability. This flexible methodology can be readily adapted to scrutinize and simulate the early propagation phases of a wide array of emerging infectious diseases. While acknowledging the paramount role of human mobility, particularly air travel, in facilitating pandemic spread, the research team also underscored the influence of a constellation of other contributing factors. These include demographic characteristics of populations, the cyclical nature of school calendars and major holiday periods that often involve increased travel and social congregation, and even prevailing weather patterns, which can impact virus survival and human behavior.
The collaborative nature of this significant research is reflected in the diverse institutional affiliations of its contributors. The study’s lead author is Renquan Zhang, affiliated with the Dalian University of Technology in Dalian, China. Additional contributions were provided by Rui Deng and Sitong Liu, also from the Dalian University of Technology; Qing Yao and Jeffrey Shaman from Columbia University; Bryan T. Grenfell from Princeton University; and Cécile Viboud from the National Institutes of Health. This international and interdisciplinary collaboration highlights the global nature of pandemic threats and the necessity of coordinated scientific efforts.
For over a decade, Professor Jeffrey Shaman and his colleagues, including Dr. Sen Pei, have been at the forefront of developing and refining sophisticated methods for tracking and simulating the transmission of highly transmissible infectious agents, including influenza viruses and SARS-CoV-2. Their ongoing work has led to the creation of advanced real-time forecasting tools that are capable of estimating the growth rate of outbreaks, predicting their likely geographical spread, and forecasting their peak intensity. These predictive capabilities are instrumental in informing public health decision-making, enabling officials to implement targeted interventions and allocate resources more effectively during critical public health emergencies. The insights gleaned from this comparative modeling study provide a crucial foundation for enhancing our collective ability to anticipate and respond to future infectious disease threats with greater speed and efficacy.