The human digestive tract harbors a vast and dynamic ecosystem, a microbial community often referred to as the gut microbiome or gut flora, which profoundly influences our overall health. This intricate consortium of microorganisms, comprising trillions of bacteria, fungi, viruses, and other microbes, is in a perpetual state of flux, shaped by an ongoing exchange of chemical signals. These exchanges occur not only among the microbial inhabitants themselves but also between the microbes and the host’s physiological environment. For this complex symbiotic relationship to function optimally, the resident bacteria must possess the ability to discern and respond to the myriad nutrients and chemical cues present in their surroundings. Despite the acknowledged significance of these microbial inhabitants, the complete spectrum of signals that bacterial sensory mechanisms can interpret remains a subject of ongoing scientific inquiry. A central, yet for a long time inadequately addressed, question has been identifying the most impactful chemical signals for the beneficial bacteria that populate our intestines.
Historically, the scientific understanding of bacterial chemosensation has been predominantly derived from studies focused on model organisms, particularly those bacteria known to cause disease. This emphasis on pathogenic microbes has resulted in a significant knowledge gap concerning the sensory capabilities of commensal bacteria – the non-pathogenic, and often beneficial, microorganisms that naturally reside within the human body. This oversight has left researchers with a limited understanding of the specific types of chemical information these helpful bacteria are actively perceiving and processing within their complex intestinal habitat.
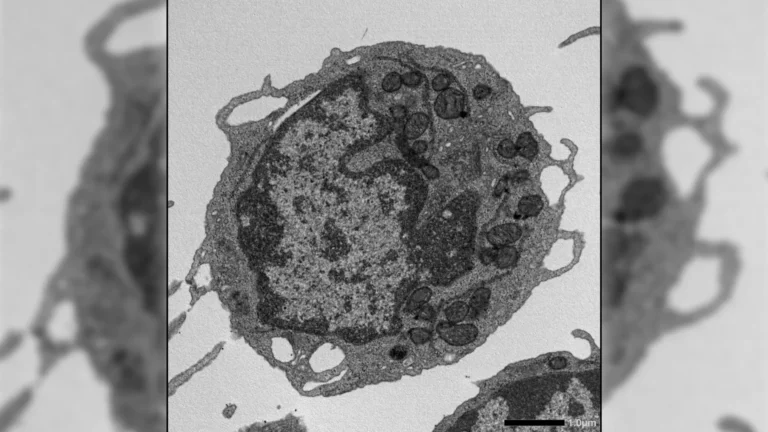
To bridge this knowledge gap, an international collaborative effort, spearheaded by Professor Victor Sourjik, was undertaken. The research team comprised scientists from distinguished institutions including the Max Planck Institute for Terrestrial Microbiology, the University of Ohio, and the Philipps-University Marburg. Their investigation zeroed in on a specific group of motile bacteria known as Clostridia, which are found in substantial numbers within the human gut and are recognized for their contributions to maintaining intestinal health.
Through meticulous laboratory investigations and sophisticated bioinformatic analyses, the researchers made a significant discovery: the sensory receptors present in the human gut microbiome exhibit an unexpectedly broad capacity to recognize a diverse range of metabolic compounds. These compounds include breakdown products originating from the digestion of carbohydrates, fats, proteins, nucleic acids (DNA), and various amines. A systematic screening process employed by the team revealed distinct patterns in these responses, demonstrating that different classes of bacterial sensory proteins exhibit specific preferences for particular categories of chemicals. This finding is crucial, as it indicates that the behavior of gut bacteria is not a haphazard reaction to their environment, but rather a finely tuned and selective response to specific metabolic indicators.
The study further elucidated the mechanisms by which motile gut bacteria navigate and locate resources. The researchers identified numerous chemical compounds, referred to as ligands, that bind to the sensory receptors governing bacterial motility. These receptors play a pivotal role in enabling motile bacteria to detect and pursue nutrients that are particularly advantageous for their proliferation and growth. The collective evidence strongly suggests that the primary impetus behind the movement of these bacteria is an active search for sustenance.
Among the extensive array of chemicals evaluated, lactic acid (lactate) and formic acid (formate) emerged as particularly potent stimuli, being recognized by a multitude of sensory receptors. This observation points to these compounds as potentially being exceptionally important nutrient sources for the resident gut bacteria. The significance of these findings is amplified by the phenomenon of cross-feeding, a process vital for the stability and health of the overall gut ecosystem. Certain bacterial species possess the capability to synthesize lactate and formate themselves, and subsequently release these compounds into their environment. These released metabolites then serve as essential food sources for other bacterial species within the same community. This intricate form of microbial cooperation helps to maintain a balanced and thriving gut environment. As Wenhao Xu, a postdoctoral researcher in Victor Sourjik’s group and the study’s lead author, explained, "These domains appear to be important for interactions between bacteria in the gut and could play a key role in the healthy human microbiome."
The research initiative also succeeded in identifying several novel groups of sensory domains, expanding the known repertoire of bacterial sensing mechanisms. These newly characterized sensors display specificity for lactate, dicarboxylic acids, uracil (a fundamental building block of RNA), and short-chain fatty acids (SCFAs) – a group of metabolites with well-established roles in host health. Furthermore, the team meticulously determined the three-dimensional crystal structure of a newly discovered dual sensor capable of responding to both uracil and acetate. This detailed structural information provided unprecedented insights into the molecular mechanisms by which these specific molecules interact with and activate the sensor. This particular sensor belongs to a larger and functionally diverse family of sensory domains found in bacteria.
An examination of the evolutionary trajectories of these uracil sensors and their related counterparts revealed a remarkable degree of evolutionary flexibility. The study demonstrated that the specificity of ligand binding can undergo relatively facile alterations over evolutionary time. This inherent adaptability is a key factor that enables bacteria to effectively adjust their sensing capabilities in response to changing environmental conditions, a critical trait for survival and colonization.
Professor Victor Sourjik summarized the project’s impact: "Our research project has significantly expanded the understanding of sensory abilities of beneficial gut bacteria. To our knowledge, this is the first systematic analysis of the sensory preferences of non-model bacteria that colonise a specific ecological niche. Looking ahead, our approach can be similarly applied to systematically investigate sensory preferences in other microbial ecosystems." This pioneering work not only deepens our comprehension of the sophisticated sensory world of gut microbes but also lays the groundwork for future investigations into the intricate workings of other microbial communities. The ability of these bacteria to precisely sense their environment, identify vital nutrients, and engage in cooperative metabolic exchanges underscores their indispensable role in maintaining human health. Understanding these complex interactions opens new avenues for therapeutic interventions and the manipulation of the gut microbiome for improved health outcomes.