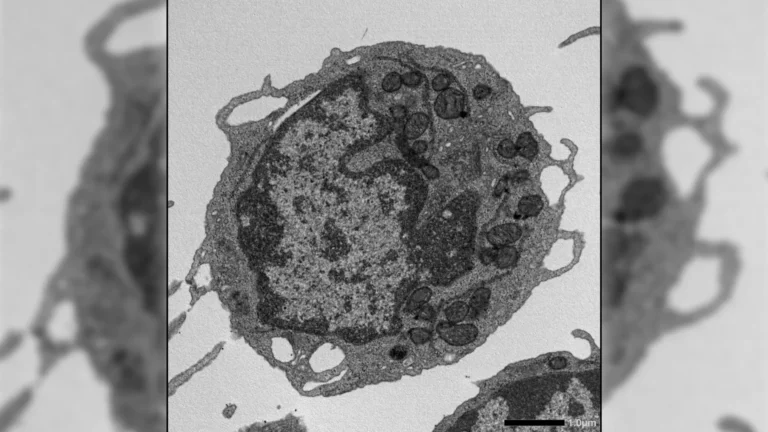
A novel bat-borne virus, provisionally named Pteropine orthoreovirus (PRV), has been detected in human patients in Bangladesh, marking a significant development in the understanding of zoonotic disease transmission. The discovery emerged from investigations into individuals who presented with symptoms suggestive of Nipah virus infection but subsequently tested negative for that particular pathogen. Researchers isolated PRV from stored throat swab samples and cultured virus from five patients, indicating that this virus should be considered in the differential diagnosis for illnesses mimicking Nipah virus in the region. This finding, published in the journal Emerging Infectious Diseases, adds to the growing list of viruses with animal origins that have been documented to infect humans in Bangladesh.
The five individuals identified with PRV infection shared a common exposure risk: they had all recently consumed raw date palm sap. This practice, prevalent in Bangladesh during the winter months, involves collecting a sweet liquid from date palm trees. Bats are known to frequent these trees, and the raw sap has previously been implicated as a primary conduit for Nipah virus transmission. Bats serve as natural reservoirs for a diverse array of zoonotic viruses, including well-known examples such as rabies, Nipah virus, Hendra virus, Marburg virus, and the precursor to SARS-CoV-1. The identification of PRV in human samples linked to this specific consumption habit underscores that the health risks associated with raw date palm sap extend beyond Nipah virus.
"Our findings highlight that the threat posed by the consumption of raw date palm sap extends beyond Nipah virus," stated Nischay Mishra, PhD, an associate professor of epidemiology at Columbia University’s Mailman School of Public Health and senior author of the study. "This also emphasizes the critical need for comprehensive surveillance programs designed to detect and manage emerging public health threats originating from bat-borne viruses."
The cases investigated occurred between December 2022 and March 2023. The patients were hospitalized with clinical presentations that included fever, vomiting, severe headaches, profound fatigue, excessive salivation, and neurological disturbances—all hallmarks of Nipah virus infection. Despite these characteristic symptoms, initial laboratory analyses, employing both polymerase chain reaction (PCR) and serological methods, failed to detect the presence of Nipah virus. To delve deeper into the etiology of these unexplained illnesses, the research team utilized high-throughput, agnostic viral capture sequencing (VCS) on the collected patient samples. This advanced molecular technique successfully identified genetic material belonging to PRV in archived throat swabs. In three of the five confirmed cases, scientists were also able to successfully cultivate the virus in laboratory settings, providing definitive confirmation of active infection.
The identification of these patients was facilitated by an ongoing Nipah virus surveillance program. This collaborative initiative is jointly managed by key public health institutions in Bangladesh, including the Institute of Epidemiology, Disease Control and Research (IEDCR) and the International Centre for Diarrheal Disease Research, Bangladesh (icddr,b), in partnership with the U.S. Centers for Disease Control and Prevention (CDC).
Viral Capture Sequencing (VCS) represents a sophisticated, patented methodology developed at Columbia University’s Center for Infection and Immunity (CII). This technology possesses the capability to screen for a comprehensive spectrum of known viral infections in vertebrate hosts, including those viruses carried by bats. The sensitivity of VCS is comparable to that of standard PCR testing, but it simultaneously searches for thousands of different viruses while also generating near-complete genomic sequences of any detected viruses. A complementary technology, Bacterial Capture Sequencing (BCS), has been developed to identify disease-causing bacteria and genes associated with antibiotic resistance. Both VCS and BCS have received approval for application in both clinical and research environments.
The five patients in this study experienced severe illness. This contrasts with some previously reported PRV infections in neighboring countries, which were often characterized by milder symptoms. This disparity suggests that less severe cases of PRV infection may be occurring in Bangladesh without being formally diagnosed, potentially leading to an underestimation of the virus’s true prevalence and impact.
"The emergence of a new zoonotic spillover event, leading to respiratory and neurological complications following the consumption of raw date palm sap, presents a significant public health concern alongside Nipah virus infection," commented Tahmina Shirin, PhD, Director of the Institute of Epidemiology, Disease Control and Research (IEDCR) and the National Influenza Centre (NIC) in Bangladesh.
Further investigations, supported by the U.S. Department of Agriculture, have aimed to pinpoint the likely source of these human infections. Researchers, including Mishra and his colleagues, have identified strains of Pteropine orthoreovirus in bats captured in proximity to the locations where the human cases occurred, specifically within the Padma River Basin. While this finding is currently unpublished, it provides crucial epidemiological evidence linking bat populations to the human infections observed.
"This research offers critical evidence connecting bat reservoirs to human transmission," stated Ariful Islam, a bat-borne disease ecologist and epidemiologist at Charles Sturt University, Australia, and a co-first author of the study. "Our ongoing efforts are focused on elucidating the precise mechanisms of spillover from bats to humans and domestic animals, as well as understanding the broader ecological dynamics of emerging bat-borne viruses within communities situated along the Padma River Basin."
The study was jointly led by Sharmin Sultana, an assistant professor of Virology and a Senior Scientific Officer at the IEDCR in Bangladesh. Additional contributions to the research came from James Ng, Sunil Kumar Dubey, Cheng Guo, and W. Ian Lipkin from the CII; Manjur Hossain Khan from IEDCR; Mohammed Ziaur Rahman and Moinuddin Satter from icddr,b; Joel M. Montgomery from the CDC’s National Center for Emerging and Zoonotic Infectious Diseases; and Lisa Hensley from the Zoonotic and Emerging Disease Research Unit at the United States Department of Agriculture. The research received funding through agreements between the United States Department of Agriculture and Columbia University.