For over a century, bacteriophages—viruses that specifically infect and destroy bacteria—have been recognized for their therapeutic potential against bacterial infections. This historical application has experienced a significant resurgence in interest as the escalating crisis of antibiotic resistance poses an increasingly formidable global health challenge. Historically, the advancement of phage-based therapies has been predominantly centered on the study and utilization of naturally occurring viral strains. This focus is largely attributable to the inherent limitations of conventional methods for modifying these viruses, which are often characterized by their slow pace, intricate complexity, and difficulties in scaling up production and application.
A groundbreaking study, recently published in the esteemed journal Proceedings of the National Academy of Sciences (PNAS), details the creation of the first fully synthetic system capable of engineering bacteriophages. This innovative approach targets Pseudomonas aeruginosa, a bacterium notorious for its high levels of antibiotic resistance and its designation as a critical threat to public health worldwide. The research was a collaborative effort between scientists at New England Biolabs (NEB) and Yale University, leveraging NEB’s proprietary High-Complexity Golden Gate Assembly (HC-GGA) platform. This advanced system empowers researchers to design and construct phages de novo from digital DNA sequence data, bypassing the necessity of relying on pre-existing viral samples.
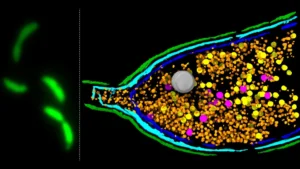
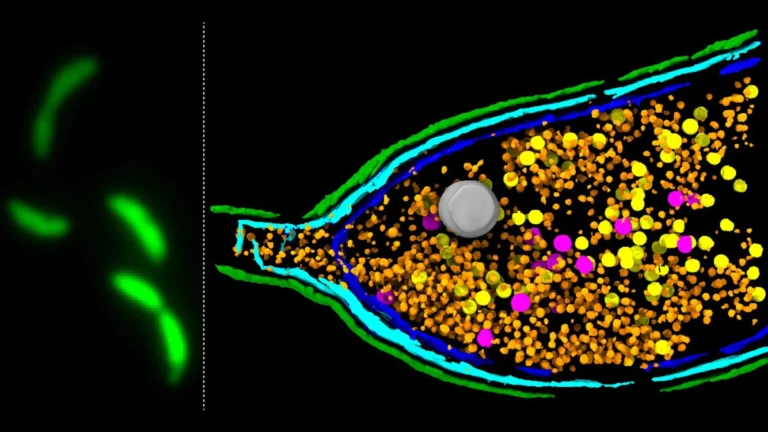
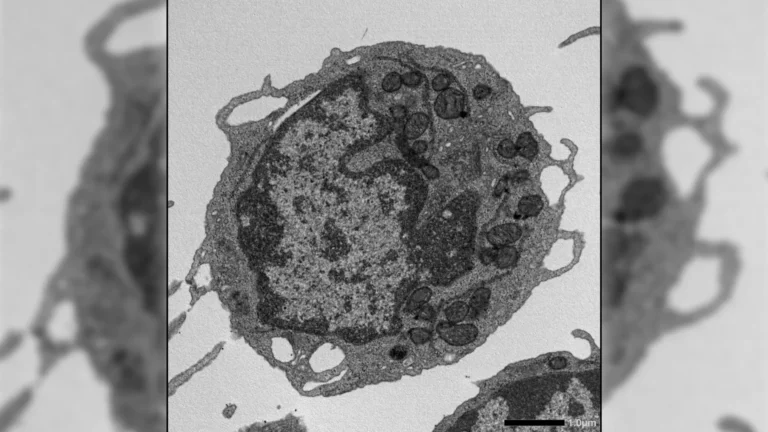
Through this novel synthetic framework, the research team successfully assembled a P. aeruginosa-targeting phage utilizing 28 distinct synthetic DNA fragments. Furthermore, they endowed the engineered virus with novel functionalities by precisely introducing point mutations and performing insertions and deletions within its genetic material. These carefully orchestrated genetic modifications enabled the researchers to, for instance, modify tail fiber genes to alter the range of bacterial species susceptible to infection by the phage. Additionally, they integrated fluorescent markers, allowing for the real-time visualization of infection processes, a significant advancement for understanding viral-bacterial interactions.
Andy Sikkema, a co-first author of the study and a Research Scientist at NEB, emphasized the transformative impact of this new methodology, stating, "Even in the most optimal scenarios, bacteriophage engineering has historically been an exceedingly labor-intensive endeavor. Researchers often dedicated their entire careers to developing intricate processes for engineering specific model bacteriophages within host bacteria. This synthetic method represents a significant technological leap forward, offering unparalleled advancements in simplicity, safety, and speed, thereby paving the way for accelerated biological discoveries and the development of new therapeutic agents."
The core of this synthetic approach lies in the ability to assemble an entire phage genome outside of a living cell. The NEB Golden Gate Assembly platform facilitates this by enabling scientists to meticulously construct the phage’s genetic blueprint using custom-synthesized DNA fragments, incorporating all intended genetic modifications during the assembly process itself. Once the complete synthetic genome is assembled, it is then introduced into a controlled, safe laboratory strain of bacteria, where it is replicated and ultimately gives rise to an active bacteriophage.
This strategic departure from traditional methods effectively circumvents numerous long-standing impediments in phage research. Conventional techniques necessitate the maintenance of physical phage stocks and the use of specialized host bacteria for propagation and manipulation. This can be particularly challenging when working with viruses that infect particularly dangerous human pathogens, where containment and specialized laboratory conditions are paramount. The newly developed synthetic method obviates the need for these demanding requirements, simultaneously eliminating the often tedious and time-consuming steps of repeated screening or sequential genetic edits performed within living host cells.
The efficacy and advantages of the Golden Gate Assembly platform in this context are multifaceted. Unlike alternative DNA assembly methodologies that typically combine fewer, but longer, DNA fragments, the Golden Gate Assembly system employs shorter, more manageable DNA segments. The production of these shorter fragments is generally more straightforward, they exhibit lower toxicity to host cells, and they are less prone to accumulating errors during synthesis. Critically, this method demonstrates remarkable robustness and compatibility with phage genomes that possess challenging genetic characteristics, such as extensive repetitive sequences or extreme guanine-cytosine (GC) content. These features often present significant obstacles for conventional DNA assembly techniques.
By streamlining the engineering process and expanding the scope of what is technically feasible, this innovative approach significantly broadens the horizon for the development of bacteriophages as highly specific and targeted therapeutic agents against the growing menace of antibiotic-resistant infections.
The genesis of this rapid synthetic phage engineering system can be attributed to a close and synergistic collaboration between the dedicated scientists at NEB and leading bacteriophage researchers at Yale University. NEB had invested years in refining the Golden Gate Assembly platform, meticulously optimizing its capacity to reliably handle and assemble large DNA targets composed of numerous individual fragments. Recognizing the profound implications of these advancements for the field of phage biology, Yale researchers proactively engaged with NEB to explore and implement more ambitious applications of the technology.
Initially, NEB scientists meticulously fine-tuned the Golden Gate Assembly method using a well-characterized model virus, the Escherichia coli phage T7, a common subject in virology research. Subsequently, collaborative teams extended the application of this refined technique to encompass non-model phages, specifically those engineered to target some of the most recalcitrant and antibiotic-resistant bacteria known to science.
This pioneering work builds upon a growing body of research showcasing the versatility of synthetic biology in phage development. For instance, a related study, also published in PNAS in November 2025, utilized the same Golden Gate approach to construct high-GC content Mycobacterium phages. This research was conducted in collaboration with the esteemed Hatfull Lab at the University of Pittsburgh and Ansa Biotechnologies, further underscoring the collaborative spirit driving innovation in this field. In another compelling application, researchers from Cornell University partnered with NEB to engineer synthetic T7 phages that function as highly sensitive biosensors. These engineered phages are capable of detecting the presence of E. coli in drinking water, as detailed in a December 2025 publication in an ACS journal, illustrating the broad applicability of synthetic phage technology beyond direct therapeutic use.
Greg Lohman, a Senior Principal Investigator at NEB and a co-author on the PNAS study, eloquently described the dynamic of innovation, stating, "My lab is dedicated to developing versatile tools, which we often refer to as ‘weird hammers,’ and then actively seeking the ‘right nails’ – the specific problems or applications where these tools can be most effectively applied. In this particular instance, the phage therapy community expressed a clear need, essentially telling us, ‘That’s precisely the hammer we have been eagerly awaiting.’" This statement highlights how the development of foundational synthetic biology tools can directly address urgent needs in critical areas of medical research and public health.