An extraordinary scientific breakthrough, detailed in the prestigious journal Science, has dramatically extended the known history of Treponema pallidum, the bacterial species responsible for a suite of debilitating infectious diseases including syphilis, yaws, and bejel. Researchers successfully reconstructed the complete genome of this ancient bacterium from human remains dating back approximately 5,500 years, discovered within the Sabana de Bogotá region of Colombia. This groundbreaking achievement, utilizing advanced paleogenomic techniques, pushes back the documented genetic presence of these pathogens in human populations by more than three millennia, fundamentally altering our understanding of their evolutionary trajectory and their long-standing association with humanity.
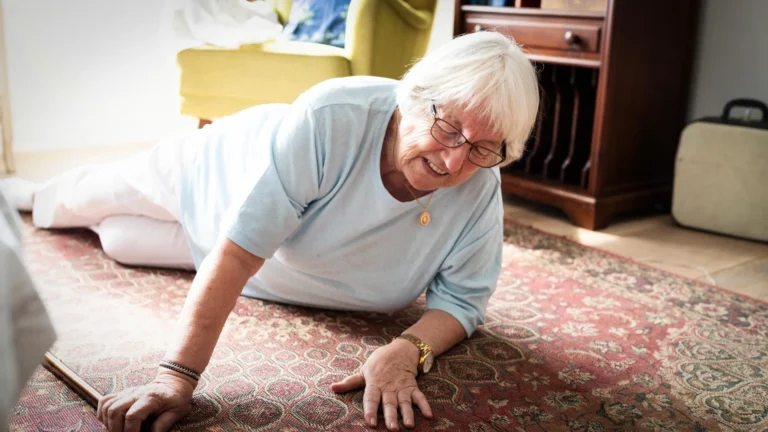
The individual whose remains yielded this crucial genetic material was unearthed from a rock shelter site near what is now modern-day Bogotá. This archaeological context places the individual within a period of significant human activity in the Americas, offering a unique window into the health challenges faced by early inhabitants. Identifying such an ancient genome provides compelling evidence that treponemal diseases were circulating across the American continents far earlier than previously established, adding substantial weight to theories proposing a deep indigenous origin for these infections in the New World.
Lars Fehren-Schmitz, a geneticist affiliated with the University of California, Santa Cruz, underscored the profound implications of this study. He articulated that "Our findings showcase the unparalleled capability of paleogenomics to contribute to our comprehension of species evolution, and to illuminate potential health risks that have confronted both historical and contemporary communities." His statement highlights not only the historical significance but also the enduring relevance of such research for public health today.
Treponemal diseases are caused by spiral-shaped bacteria belonging to the genus Treponema. The species Treponema pallidum is particularly notorious, comprising several closely related subspecies, each responsible for a distinct clinical syndrome. Treponema pallidum subspecies pallidum causes syphilis, a sexually transmitted infection with widespread systemic effects. Treponema pallidum subspecies pertenue is the causative agent of yaws, a chronic skin, bone, and cartilage infection typically found in humid tropical regions and transmitted through skin-to-skin contact. Bejel, or endemic syphilis, caused by Treponema pallidum subspecies endemicum, is another non-venereal treponemal disease affecting skin, bones, and mucous membranes, prevalent in arid regions and spread through close contact. A fourth treponemal disease, pinta, primarily affecting the skin, is caused by Treponema carateum (sometimes classified as Treponema pallidum subspecies carateum). Intriguingly, a full genomic sequence for the pathogen responsible for pinta has yet to be recovered, leaving considerable gaps in our knowledge regarding its evolutionary relationships and precise classification within the treponemal family.
Despite the remarkable genetic similarity among these various subspecies, the precise timing and mechanisms through which these distinct disease forms emerged remain largely enigmatic. While physical evidence of past infections can sometimes be discerned through macroscopic examination of skeletal remains—manifesting as characteristic lesions or bone deformities—the story told by ancient DNA often proves far more intricate and revealing. There frequently exists a considerable disparity between the observable pathology on ancient bones and the nuanced insights that ancient genomic analysis can provide regarding the evolution of pathogens.
The research team confirmed that the ancient DNA isolated from the Colombian remains unequivocally belonged to the species Treponema pallidum. However, a critical finding was that this ancient genome did not genetically align with any of the currently recognized forms responsible for human disease today. Although undeniably a close relative to modern strains, this ancient genetic lineage appears to have diverged very early in the bacterium’s complex evolutionary history. This discovery points to the existence of a "lost lineage" of Treponema pallidum, one that branched off long before the emergence of the subspecies we are familiar with today.
Anna-Sapfo Malaspinas, a group leader at the SIB Swiss Institute of Bioinformatics and associated with the University of Lausanne, offered a compelling hypothesis regarding this unique finding. She speculated, "One possibility is that we have unearthed an ancient variant of the pathogen that causes pinta, a disease about which we possess limited information, but which is known to be endemic across Central and South America and typically produces localized skin manifestations." She cautioned, however, that while this remains an intriguing lead worthy of further exploration, definitive proof is currently elusive.
Genetic analyses allowed scientists to estimate the divergence time for this ancient strain, placing its separation from other T. pallidum lineages at approximately 13,700 years ago. This timeframe stands in stark contrast to the estimated divergence of the three modern subspecies (syphilis, yaws, bejel), which appear to have branched much more recently, around 6,000 years ago. These chronological data provide robust support for earlier research indicating a much greater diversity among treponemal pathogens in the distant past than previously appreciated.
Elizabeth Nelson, a molecular anthropologist and paleopathologist at Southern Methodist University (SMU), emphasized the broader implications of these findings. She noted, "Current genomic evidence, coupled with the genome we present here, does not definitively resolve the enduring debate surrounding the ultimate geographical origins of the disease syndromes themselves. Nevertheless, it unequivocally demonstrates a profound and extensive evolutionary history of treponemal pathogens, which were already diversifying within the Americas millennia before our prior knowledge suggested." This statement highlights the intricate relationship between pathogen evolution and human migration patterns, particularly in the context of the contentious "Out of America" versus "Out of Africa" debate concerning the origins of syphilis.
Tracing the evolutionary origins of treponemal diseases presents a particularly formidable challenge due to the remarkable genetic similarity shared by the causative bacteria. Paradoxically, despite this genetic uniformity, these pathogens employ diverse transmission routes and can induce a wide spectrum of clinical symptoms, rendering their evolutionary pathways exceedingly difficult to unravel. Davide Bozzi, a researcher at the University of Lausanne and the SIB Swiss Institute of Bioinformatics, underscored the magnitude of this discovery, stating, "Our findings extend the established association of T. pallidum with human populations by thousands of years, potentially reaching back more than 10,000 years into the Late Pleistocene epoch."
This pivotal discovery is the culmination of extensive, long-term archaeological and genetic investigations conducted at the Tequendama 1 site. Prior studies led by archaeologist Miguel Delgado of the Universidad Nacional de La Plata in Argentina, in collaboration with Fehren-Schmitz, had already provided detailed contextual information about the skeletal remains themselves, setting the stage for this unprecedented genomic analysis.
Remarkably, the detection of the pathogen was not an intentional initial objective of the research. The team was primarily engaged in sequencing the ancient individual’s DNA to investigate early human population history in the region. This endeavor generated an astonishing volume of genetic data—approximately 1.5 billion fragments—a quantity far exceeding typical outputs for ancient DNA studies. During a routine screening process, research teams operating independently at the University of California, Santa Cruz, and the University of Lausanne, both detected faint but unmistakable traces of T. pallidum DNA. Recognizing the immense potential of this unexpected find, they decided to consolidate their efforts and collaborate on a joint investigation.
Despite the fact that bacterial DNA constituted only a minuscule fraction of the total genetic material recovered, the sheer depth of sequencing achieved enabled the team to reconstruct the pathogen’s complete genome. This was accomplished without the necessity of specialized enrichment techniques, which are often required to boost the concentration of target DNA in ancient samples. This speaks volumes about the power of ultra-deep sequencing in unlocking hidden genomic information.
A crucial aspect of this discovery lies in the fact that the diseases caused by T. pallidum (such as bejel, yaws, and syphilis) can manifest as distinctive lesions on bones, but only under specific conditions and not in every infected individual. Historically, most ancient genomes of this bacterium have been successfully recovered from teeth or skeletal elements that presented clear, visible signs of disease. In this particular instance, however, the ancient skeleton showed no discernible evidence of infection. Furthermore, researchers sampled a tibia, or shin bone, a skeletal element not conventionally favored for ancient DNA studies due to its typically lower preservation quality compared to denser bones or teeth. The success of this unconventional approach strongly suggests that even bones lacking obvious pathological markers can, under favorable conditions, preserve invaluable genetic information, opening new avenues for paleogenomic research.
Understanding the historical trajectories of infectious diseases holds profound relevance for contemporary global health challenges. By meticulously charting how pathogens emerged, evolved, and adapted in the past, scientists aspire to develop a more robust predictive capacity for anticipating how these microbial threats might transform in the future. This invaluable knowledge could serve as a critical foundation for modern societies to proactively prepare for and mitigate potential future health crises, ranging from emerging infectious diseases to the challenges of antibiotic resistance.
Prior to the publication of their groundbreaking results, the international research team proactively shared their findings with communities in Colombia. This engagement recognized the profound cultural and medical historical significance of the discovery for the country. They consulted extensively with local scholars, students, and both Indigenous and non-Indigenous community members, fostering dialogue and understanding through a series of presentations and interviews with key stakeholders. All requisite permits for the export and study of the ancient remains were meticulously obtained, adhering to international ethical guidelines.
Miguel Delgado underscored the imperative nature of this community engagement process. He stated, "This process was absolutely essential because the findings are intricately woven into Colombia’s medical and cultural heritage." He further elaborated that "Engaging scholars, students, and both Indigenous and non-Indigenous community members ensures that the results are communicated ethically and interpreted in genuine partnership with local communities. This collaborative approach fosters trust, supports the responsible stewardship of sensitive scientific discoveries, and ultimately reinforces local ownership of this crucial knowledge."
This monumental research was a testament to international collaboration, involving a diverse group of experts. In addition to Nelson, Bozzi, Malaspinas, Delgado, and Fehren-Schmitz, the study was co-led by Nasreen Broomandkhoshbacht, now affiliated with the University of Vermont. The broader research collective included Kalina Kassadjikova of the University of California, Santa Cruz; Jane Buikstra of Arizona State University; Carlos Eduardo G. Amorim of California State University, Northridge; Melissa Estrada Pratt of the Instituto Colombiano de Antropología e Historia in Bogotá, Colombia; Gilbert Greub of the University of Lausanne and Lausanne University Hospital in Switzerland; Nicolas Rascovan of the Institut Pasteur in Paris; and David Šmajs of Masaryk University in the Czech Republic. Their combined expertise made possible this extraordinary glimpse into the deep history of human-pathogen interactions.