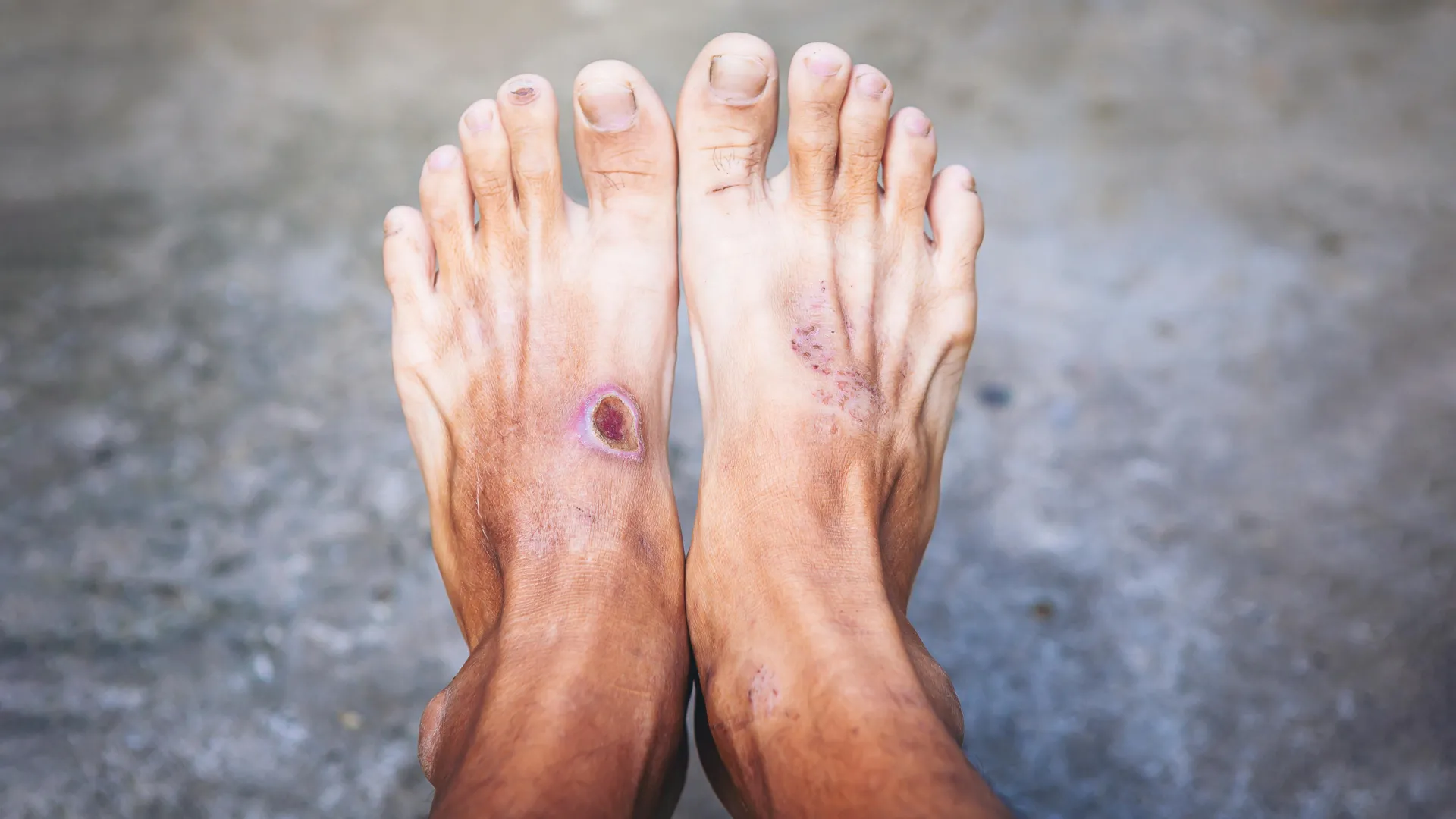
Diabetes mellitus, a chronic metabolic disorder affecting hundreds of millions worldwide, carries with it a spectrum of debilitating complications. Among the most severe and life-altering are diabetic foot infections (DFIs), which represent a significant global health challenge. These complex, chronic wounds are not merely superficial injuries; they are a leading precursor to non-traumatic lower-limb amputations, profoundly impacting patient quality of life, increasing healthcare costs, and often resulting in severe disability or even mortality. For decades, clinicians have grappled with the polymicrobial nature of these infections, where a diverse array of bacteria can colonize and exacerbate the wounds. However, a recent landmark study, spearheaded by researchers at King’s College London in collaboration with the University of Westminster, has cast a revealing light on one particularly prevalent yet historically under-characterized pathogen in this context: Escherichia coli. This extensive genomic analysis offers unprecedented insights into the genetic makeup, diversity, and antimicrobial resistance profiles of E. coli strains isolated from diabetic foot ulcers across ten countries, fundamentally reshaping our understanding of this microscopic threat.
The investigation, published in the peer-reviewed journal Microbiology Spectrum, stands out as the first comprehensive, large-scale genomic examination of E. coli directly sampled from infected diabetic foot ulcers on an international scale. By delving into the complete DNA sequences of these bacterial strains, the research team sought to unravel the underlying biological mechanisms that render some DFIs exceptionally challenging to treat, often leading to prolonged hospitalization, sepsis, and the ultimate devastating outcome of amputation. This deep genomic dive transcends traditional microbiological techniques, moving beyond simple identification to expose the intricate genetic blueprints that govern bacterial behavior, adaptability, and pathogenicity within the unique environment of a diabetic wound.
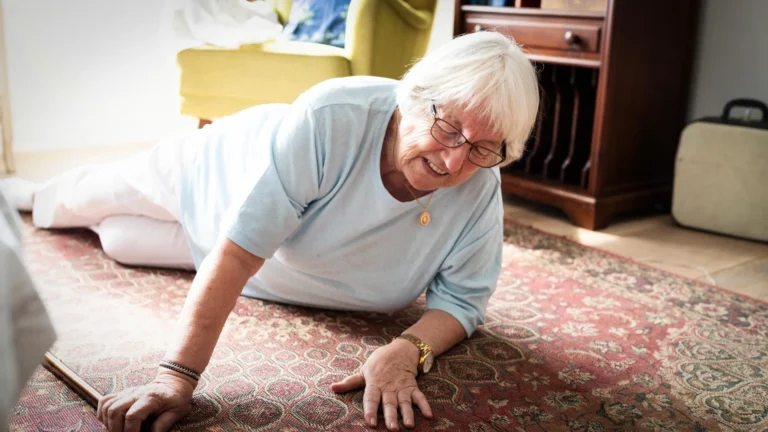
Diabetic foot infections present a formidable clinical conundrum. The underlying pathology of diabetes, including peripheral neuropathy (nerve damage), peripheral artery disease (poor blood circulation), and impaired immune function, creates an ideal environment for microbial proliferation and deep tissue invasion. The compromised sensation means minor injuries can go unnoticed, while poor circulation hinders healing and antibiotic delivery. Coupled with a weakened immune response, these factors make DFIs particularly susceptible to severe, rapidly progressing infections. While the presence of various bacterial species in these wounds has long been acknowledged, the specific role and characteristics of E. coli have often remained somewhat opaque. Despite its frequent isolation from DFI samples, E. coli has not received the same level of in-depth study as some other common DFI pathogens, such as Staphylococcus aureus or Pseudomonas aeruginosa. This knowledge gap has impeded the development of targeted diagnostic tools and therapeutic strategies.
To bridge this critical informational divide, the research consortium embarked on an ambitious global sampling initiative. They meticulously collected and analyzed 42 distinct E. coli strains, each derived from an infected diabetic foot ulcer. The geographical breadth of this collection was truly global, encompassing samples from patients spanning continents: Nigeria, the United Kingdom, Ghana, Sweden, Malaysia, China, South Korea, Brazil, India, and the United States. This international representation was crucial for identifying universal patterns in E. coli‘s behavior in DFIs, as well as potential regional variations. The core of their methodology involved whole-genome sequencing (WGS) for each strain. WGS provides a complete genetic blueprint, allowing scientists to meticulously compare minute differences between strains, identify genes conferring resistance to antibiotics, and pinpoint specific biological traits, known as virulence factors, that contribute to the severity and progression of the disease. This high-resolution genetic mapping offered an unprecedented opportunity to dissect the evolutionary relationships and functional capabilities of E. coli in this clinical context.
One of the most profound discoveries emerging from this extensive genomic investigation was the remarkable genetic diversity among the E. coli strains. Contrary to a potential expectation of a few dominant, highly adapted lineages, the findings unequivocally demonstrated that E. coli strains implicated in diabetic foot infections belong to a multitude of different genetic groups. They possessed a vast array of genes associated with disease causation (virulence) and resistance to antimicrobial agents. This striking heterogeneity disproved the notion that DFIs are primarily caused by a singular, specialized type of E. coli. Instead, the evidence suggested that numerous distinct and often unrelated E. coli lineages have independently evolved or acquired traits enabling them to thrive within the challenging and complex milieu of the diabetic foot environment. This revelation underscores the intricate nature of DFI pathogenesis and highlights why a one-size-fits-all approach to treatment often falls short. By understanding the genomic relatedness of these strains and their specific arsenals of resistance mechanisms and virulence factors – the molecular tools that enhance a microbe’s ability to infect and cause harm – the study provides crucial mechanistic insights into why some infections are recalcitrant to standard therapies or progress with alarming rapidity.
A particularly alarming finding from the study centered on the prevalence of antimicrobial resistance (AMR). The analysis revealed that approximately 8% of the E. coli strains identified were classified as either multidrug-resistant (MDR) or extensively drug-resistant (XDR). Multidrug-resistant bacteria are those that exhibit resistance to at least one agent in three or more antimicrobial categories, while extensively drug-resistant bacteria are resistant to nearly all available antibiotic classes, leaving very few, if any, effective treatment options. This statistic, though seemingly small, carries immense clinical implications. The presence of MDR and XDR E. coli significantly complicates patient management, often necessitating the use of last-resort antibiotics, prolonging hospital stays, increasing healthcare costs, and drastically elevating the risk of treatment failure, severe complications, and ultimately, limb loss. This finding resonates with the broader global health crisis of antimicrobial resistance, emphasizing its critical impact on specific, vulnerable patient populations like those with diabetic foot ulcers.
Dr. Vincenzo Torraca, a Lecturer in Infectious Disease at King’s College London and a senior author of this pivotal study, underscored the transformative potential of these genomic insights. He stated, "Deciphering the genomic architecture of these bacteria is an indispensable stride towards refining diagnostic precision and enabling more personalized therapeutic interventions for individuals living with diabetes. By accurately identifying the predominant E. coli strains and anticipating their likely resistance profiles, clinicians can transition from empirical, broad-spectrum antibiotic regimens to highly targeted therapies. This strategic shift is crucial for mitigating prolonged infection courses, reducing hospital admissions, and ultimately diminishing the devastating risk of amputation." His comments highlight the promise of precision medicine in infectious disease, where genetic information guides treatment decisions.
Echoing the far-reaching implications of the research, Victor Ajumobi, a second-year PhD student at King’s College London and the University of Westminster, who served as the paper’s first author, emphasized its particular relevance for underserved communities. "This critical information will prove especially invaluable in low-resource healthcare settings," Ajumobi noted, "where E. coli infections within diabetic foot ulcers are often more prevalent, and where access to sophisticated, rapid diagnostic tools for antimicrobial resistance is frequently limited or non-existent." This perspective highlights the study’s potential to inform public health strategies and clinical guidelines in regions where the burden of DFIs and AMR is often highest, and where resources for advanced diagnostics are scarce. Understanding common resistance patterns in these areas could lead to more effective empiric treatment guidelines, saving limbs and lives.
Looking ahead, the research team is not resting on its laurels. Their immediate plans involve delving deeper into the functional consequences of the specific virulence factors identified through the genomic analysis. Many of the bacterial samples were found to harbor genes that equip E. coli with capabilities such as enhanced adherence to human tissues or sophisticated mechanisms to evade the host immune system. Investigating how these particular traits manifest and operate within the complex environment of diabetic foot wounds could unlock novel therapeutic targets. This future research could pave the way for the development of innovative treatment strategies that go beyond traditional antibiotics, potentially including anti-virulence drugs that disarm the bacteria rather than merely killing them. Such advancements could support the creation of more effective, less resistance-prone therapies, ultimately improving patient outcomes and significantly reducing the global burden of diabetic foot amputations. This comprehensive genomic investigation marks a pivotal moment in the fight against diabetic foot infections, providing a robust scientific foundation for future clinical and public health interventions.