The escalating global threat posed by antibiotic-resistant bacteria, commonly known as superbugs, has spurred an intensive scientific quest for innovative therapeutic avenues, with recent breakthroughs illuminating a critical vulnerability within bacterial cell wall construction. A team of researchers has meticulously detailed how specific viral proteins, evolved independently, converge on a single, indispensable bacterial protein, effectively disabling the machinery responsible for building the bacterial cell’s protective outer layer. This discovery, published in the prestigious journal Nature, not only deepens our understanding of bacterial physiology but also pinpoints a highly promising target for the development of next-generation antibiotics.
The urgency driving this research cannot be overstated. Bacteria possess an extraordinary capacity for rapid adaptation, a trait that has led to the alarming emergence of strains impervious to virtually all existing antimicrobial agents. This evolutionary prowess transforms a once-manageable public health concern into a critical crisis, with tens of thousands of fatalities annually in the United States alone attributed to infections that defy conventional treatment, a number that continues its upward trajectory. The dwindling efficacy of current pharmaceuticals necessitates a radical shift in strategy, compelling scientists to explore entirely new molecular mechanisms that bacteria rely upon for survival.
A long-standing focus in antimicrobial research has been the intricate pathway bacteria employ to synthesize peptidoglycan, a complex polymer that forms the rigid, essential scaffold of their cell walls. This process, known as peptidoglycan biosynthesis, represents a particularly attractive target for drug development due to its exclusive presence in bacterial cells, rendering it absent in human tissues. Consequently, interventions aimed at disrupting this pathway are less likely to induce collateral damage to host cells, offering a safer therapeutic profile. While established antibiotics, such as penicillin and its derivatives, already interfere with later stages of peptidoglycan production, the search continues for agents that can disrupt earlier, perhaps even more fundamental, steps in this vital process.
Central to the transport of peptidoglycan building blocks across the bacterial inner membrane are three key proteins: MraY, MurG, and MurJ. These proteins act as molecular chaperones, facilitating the movement of the essential components required for the assembly of the cell wall structure on the exterior of the cell membrane. The failure of any of these proteins to perform their function would halt peptidoglycan production, leading to the demise of the bacterium. Therefore, these proteins represent highly desirable targets for therapeutic intervention. Despite considerable progress in understanding their general roles, specific mechanistic details regarding their operation have remained elusive, hindering the development of direct inhibitors. While no currently approved drugs directly target these three proteins, the potential for identifying small molecules, either naturally derived or synthetically engineered, that can inhibit their activity is significant.
Intriguingly, the natural world has already provided insights into how these bacterial mechanisms can be subverted. Bacteriophages, viruses that specifically infect bacteria, must navigate the peptidoglycan layer to complete their life cycle, either by entering the host cell or, more critically for their propagation, by escaping after replication. To exit a bacterial cell, phages must breach this formidable peptidoglycan barrier, which functions akin to a molecular armor. Failure to do so would trap them within the host.
The research team, specifically focusing on bacteriophages with compact genomes, such as those containing single-stranded DNA or RNA, has investigated their sophisticated strategies for bacterial lysis. These minimalist viruses often employ specialized protein components, termed single-gene lysis proteins (Sgls), to orchestrate bacterial demise. The current study zeroes in on Sgls that specifically target MurJ, one of the crucial proteins involved in cell wall synthesis.
MurJ functions as a flippase, a type of transporter protein that moves molecules across a membrane. In the context of peptidoglycan synthesis, MurJ is responsible for translocating peptidoglycan precursors from the interior of the bacterial cell to the periplasmic space, where they can be incorporated into the growing cell wall. Previous research had established that two distinct Sgls, identified as SglM and SglPP7, exert their antibacterial effect by inhibiting MurJ.
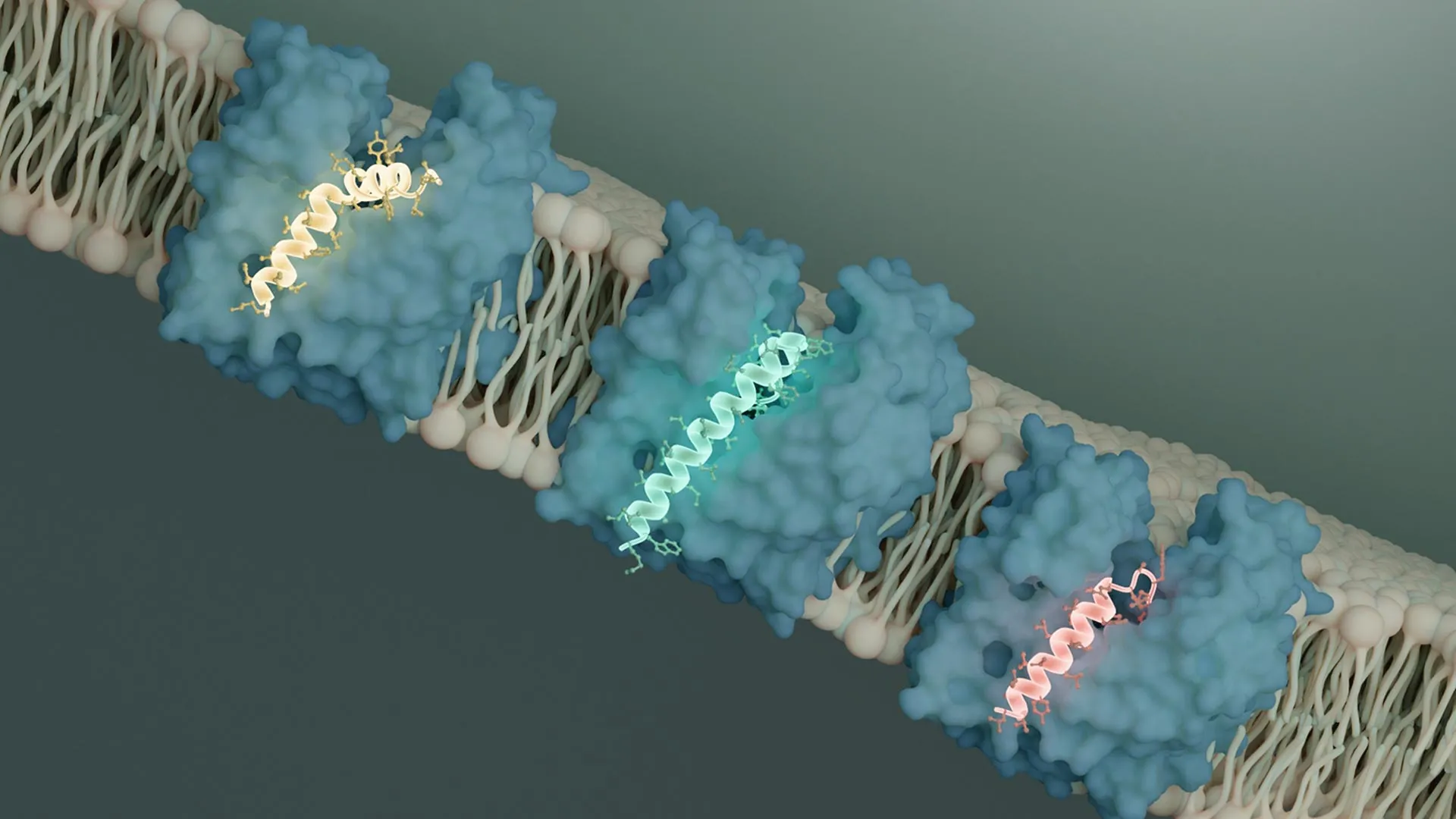
Employing cutting-edge cryo-electron microscopy (cryo-EM) techniques at Caltech’s Beckman Institute Biological and Cryogenic Transmission Electron Microscopy (Cryo-EM) Resource Center, the researchers were able to visualize the precise molecular interactions between these viral proteins and MurJ. Their findings revealed that both SglM and SglPP7 bind to a specific groove on the MurJ protein. This binding event physically obstructs MurJ’s conformational change, a critical step required for it to transport its molecular cargo across the membrane.
More significantly, the structural analysis indicated that both SglM and SglPP7 bind to MurJ when the protein is in an outward-facing conformation. This orientation is crucial because it means the protein’s active site is exposed towards the exterior of the bacterial cell, potentially making it more accessible to therapeutic agents compared to an inward-facing configuration. This observation is particularly encouraging for drug development, as it suggests a favorable structural characteristic for targeting MurJ.
The convergence of two evolutionarily distant viral proteins on the same functional site of MurJ in a remarkably similar manner represents a striking example of convergent evolution. This phenomenon, where unrelated organisms independently evolve similar traits or solutions to similar environmental challenges, underscores the inherent biological significance and vulnerability of the MurJ protein. The researchers postulate that given the rapid evolutionary pace of bacteriophages, it is highly probable that numerous other phages harbor analogous Sgls capable of targeting MurJ. The relative ease of isolating phages and sequencing their genomes offers a vast reservoir for discovering new biological insights and, critically, novel antibiotic targets.
Further reinforcing this hypothesis, the research team analyzed another phage genome, identifying a novel Sgl, designated SglCJ3, derived from a phage predicted to be named Changjiang3. Through cryo-EM analysis, they determined the structure of SglCJ3 in complex with MurJ, confirming that this third, independently evolved viral protein also functions by locking MurJ into the same outward-facing conformation, thereby inhibiting its transport activity.
The consistent observation of distinct viral peptides converging on MurJ through similar inhibitory mechanisms provides compelling evidence that evolution itself has identified this protein as a prime target for bacterial disruption. This discovery strongly advocates for following nature’s lead by developing therapeutics that specifically target MurJ. The study serves as a powerful testament to the capacity of fundamental biological research to provide solutions to pressing medical challenges. The ongoing path forward involves leveraging the discovery of Sgls and translating these foundational scientific concepts into tangible therapeutic realities, a process that will likely require continued support and collaboration.
The comprehensive findings of this study are detailed in the Nature paper titled "Convergent MurJ flippase inhibition by phage lysis proteins." The research was a collaborative effort involving Yancheng Evelyn Li and Bil Clemons from Caltech, alongside Grace F. Baron, also a graduate student at Caltech, and contributions from Francesca S. Antillon, Karthik Chamakura, and Ry Young at Texas A&M University. Funding for this groundbreaking research was provided by significant grants from the Chan Zuckerberg Initiative, the National Institutes of Health, the G. Harold and Leila Y. Mathers Foundation, and the Center for Phage Technology at Texas A&M, a joint initiative supported by Texas A&M AgriLife.