In a significant advancement poised to reshape the landscape of medical diagnostics and industrial biotechnology, an international consortium of scientists has unveiled a novel class of DNA-binding proteins. These remarkable molecules, previously undocumented, originate from some of our planet’s most inhospitable locales and possess an extraordinary resilience, enabling them to dramatically enhance the speed and sensitivity of critical diagnostic tools used in the detection of infectious diseases. This breakthrough not only underscores the immense, untapped biological wealth of extreme environments but also offers profound implications for the burgeoning field of artificial intelligence in protein engineering.
The collaborative research initiative was primarily spearheaded by experts at Durham University, bringing together diverse scientific acumen from partner institutions situated in Iceland, Norway, and Poland. Their ambitious investigation involved a meticulous examination of genetic material meticulously harvested from two distinct types of extreme habitats: the geothermally active volcanic lakes nestled within Iceland’s rugged terrain and the abyssal hydrothermal vents submerged more than two kilometers beneath the formidable North Atlantic Ocean. These environments are characterized by conditions – such as searing temperatures, crushing pressures, and highly acidic or saline waters – that would be lethal to most life forms, yet they foster unique microbial ecosystems adapted to thrive under such duress.
The scientific quest for novel biomolecules often leads researchers to the farthest corners of the Earth, driven by the understanding that nature remains the most prolific and sophisticated chemist. While countless enzymes and proteins play indispensable roles in biological processes, a vast majority still await discovery, particularly those adapted to unusual conditions. To unearth these hidden treasures, the research team embarked on an intensive bioprospecting endeavor. They leveraged state-of-the-art next-generation DNA sequencing technologies to scrutinize massive bioinformatics repositories, which house millions upon millions of potential protein sequences. This advanced computational approach allowed them to sift through an ocean of genetic data, identifying patterns indicative of unique protein functions.
Through this extensive bioinformatic analysis, the team successfully pinpointed a collection of previously uncharacterized proteins exhibiting a distinct affinity for binding to single-stranded DNA. What immediately set these newly identified proteins apart was their inherent stability under a spectrum of harsh environmental stressors. Laboratory tests confirmed their exceptional fortitude, demonstrating robust functionality even when exposed to extreme thermal conditions, highly acidic or alkaline pH levels, and environments saturated with high concentrations of salt. Such intrinsic durability is a rare and highly prized attribute in the world of molecular biology and biotechnology, promising utility in demanding applications where conventional enzymes often falter.
Following their initial identification, the structural and functional properties of these robust proteins were subjected to rigorous scrutiny using an array of advanced laboratory methodologies. Scientists confirmed their exceptional durability and noted their remarkable thermal stability, a characteristic that renders them particularly well-suited for a wide spectrum of biotechnological and biomedical applications. Enzymes that can withstand high temperatures, for instance, are invaluable in industrial processes and diagnostic assays that often require elevated reaction temperatures or prolonged shelf life under varying conditions.
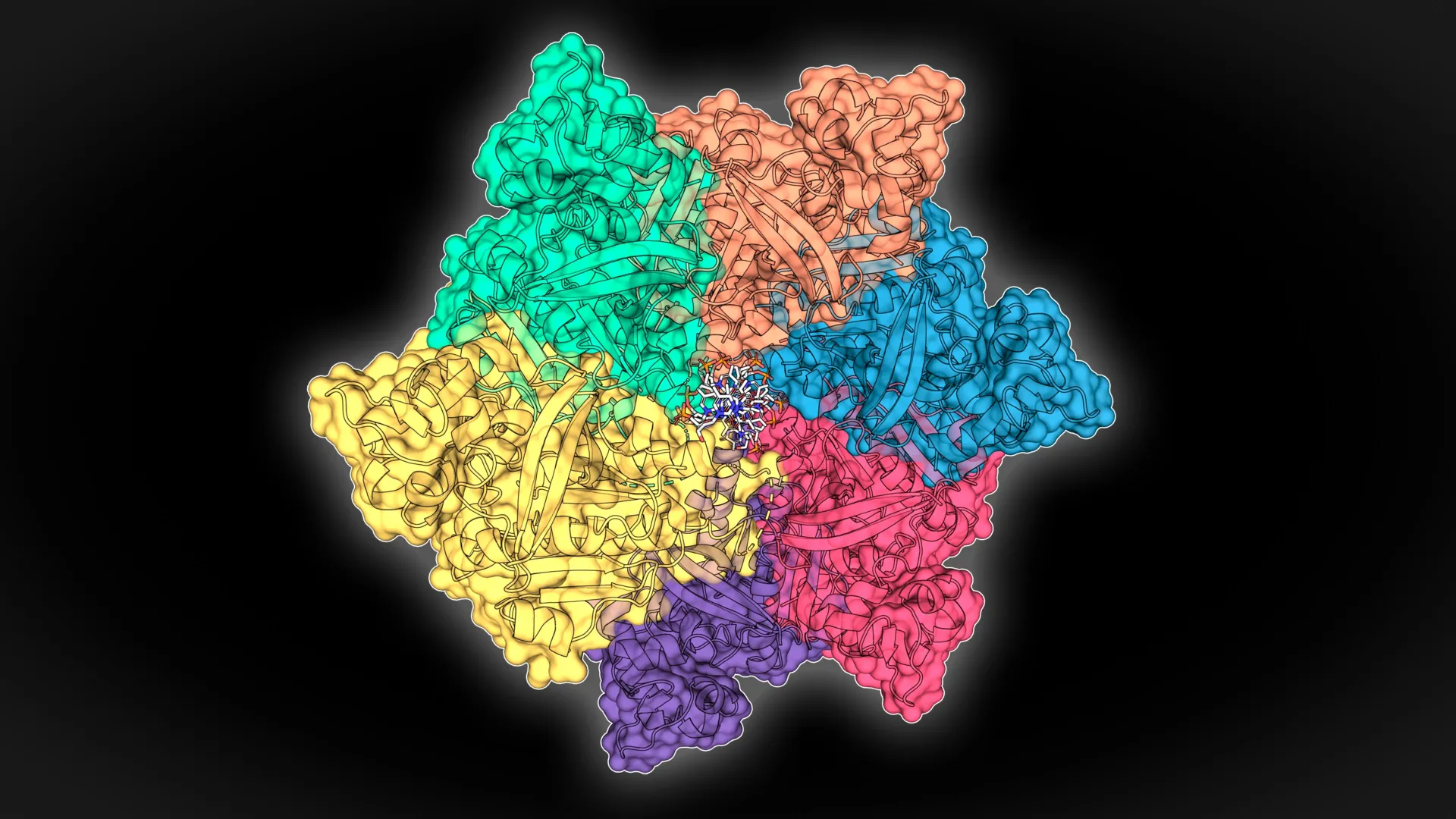
Furthermore, the research team meticulously determined the three-dimensional atomic structures of these proteins at high resolution. This detailed structural information is paramount; it offers critical insights into the precise mechanisms by which these molecules exert their functions. Understanding a protein’s architecture at an atomic level is akin to possessing the blueprint of a complex machine, revealing how its various components interact to perform a specific task. Such structural clarity not only deepens our fundamental understanding of these unique biomolecules but also provides a foundational platform for future advancements, including the rational design and engineering of proteins with enhanced or entirely new functionalities. This structural knowledge is a cornerstone for directed evolution and synthetic biology efforts aimed at optimizing protein performance for specific industrial or medical needs.
One of the most immediate and impactful applications of these newly discovered DNA-binding proteins lies in their capacity to significantly improve rapid diagnostic tests. Specifically, one of the isolated proteins demonstrated a remarkable ability to enhance loop-mediated isothermal amplification (LAMP) assays. LAMP technology represents a cornerstone of point-of-care diagnostics, offering a relatively simple, cost-effective, and rapid method for detecting genetic material from a wide array of pathogens, including viruses, bacteria, and parasites. Unlike more complex PCR-based methods, LAMP tests do not necessitate sophisticated thermal cyclers, making them ideal for deployment in resource-limited settings, remote clinics, or during outbreaks where swift, decentralized testing is crucial.
The integration of the novel extremophile protein into existing LAMP test protocols yielded tangible and impressive improvements. The assays exhibited a noticeable acceleration in their reaction times, meaning results could be obtained more quickly. Crucially, the tests also became considerably more sensitive, enabling the detection of lower concentrations of target genetic material. This dual enhancement—increased speed and heightened sensitivity—is a game-changer for diagnostic accuracy and efficiency. For example, the improved LAMP tests proved more effective at detecting viral RNA from formidable pathogens like SARS-CoV-2, the virus responsible for COVID-19, as well as DNA from various other infectious agents. The implications for public health are profound, offering the potential for earlier diagnosis, more effective disease surveillance, and swifter public health interventions, particularly in the face of emerging pandemics or widespread outbreaks.
Beyond their immediate utility in diagnostics, these findings resonate across broader scientific and economic landscapes, particularly within the biotechnology sector and the rapidly evolving field of AI-driven protein design. The global biotechnology industry is in a perpetual quest for enzymes and proteins that can perform reliably and efficiently under challenging industrial conditions, such as those involving high temperatures, extreme pH, or the presence of harsh chemicals. Proteins naturally adapted to thrive in environments like volcanic hot springs or deep-sea hydrothermal vents are inherently robust, making them exceptionally promising candidates for a multitude of industrial applications, ranging from biofuel production and bioremediation to food processing and pharmaceutical manufacturing. Their intrinsic stability translates directly into cost savings and improved process efficiency for companies operating in these demanding areas.
Furthermore, these discoveries contribute significantly to the ongoing advancements in artificial intelligence systems dedicated to protein structure prediction and rational protein design. AI models, particularly those employing machine learning and deep learning algorithms, thrive on large, diverse, and high-quality datasets. Each newly characterized protein, especially one with unique properties derived from an extreme environment, provides invaluable training data for these AI systems. By expanding the repertoire of known protein structures and functions, researchers can refine AI models, enabling them to more accurately predict the structures of unknown proteins and, crucially, to design novel proteins with tailor-made functionalities. This synergistic relationship between biological discovery and computational intelligence promises to unlock unprecedented capabilities in drug discovery, material science, and enzyme engineering, driving forward the broader bioeconomy. As Professor Ehmke Pohl of Durham University, the lead investigator of the study, articulated, this work not only holds significant value for the bioeconomy but also lays fundamental groundwork for all AI methodologies in protein structure prediction and design.
The journey of discovery for these remarkable proteins is far from over. The research team is actively pursuing further investigations into additional DNA-binding proteins, with several promising candidates already identified for future study. Concurrent efforts are focused on engineering improved variants of the discovered proteins, aiming to optimize their performance and stability even further for specific applications. A key translational objective involves designing novel LAMP tests specifically targeting neglected tropical diseases, such as leishmaniasis and Chagas disease. These debilitating conditions disproportionately affect impoverished populations and often lack adequate diagnostic tools, highlighting the potential for these new proteins to address critical global health inequities. This specialized work is being conducted in close collaboration with researchers in Durham University’s Biosciences Department, integrating expertise across disciplines.
Moreover, the commercialization potential of these groundbreaking discoveries is being actively explored through a strategic partnership with ArcticZymes, a Norwegian biotechnology company. This collaboration aims to bridge the gap between fundamental scientific research and practical market applications, ensuring that the benefits of these robust proteins can be translated into tangible products and solutions for both medical diagnostics and industrial biotechnology. This comprehensive approach, spanning from deep-sea exploration to advanced AI applications and commercial development, exemplifies the multidisciplinary nature of modern scientific innovation and its profound potential to address some of humanity’s most pressing challenges.