The analytical technique of mass spectrometry boasts a history stretching back to the early 20th century, establishing itself as an indispensable tool across a myriad of scientific disciplines. From identifying the intricate components of a cell to ensuring the purity of pharmaceutical compounds, its fundamental ability to discern the precise molecular makeup of a sample and quantify its constituents is unparalleled. Despite its profound impact and widespread application, a persistent operational constraint has long limited its full potential: the vast majority of contemporary mass spectrometers are designed to process molecules sequentially, either one at a time or in exceedingly small batches. This inherent limitation translates into significant challenges, including protracted analysis times, elevated operational costs, and, critically, an increased likelihood of overlooking rare yet highly significant molecules that are often obscured by more abundant species within complex matrices.
A recent breakthrough from researchers at The Rockefeller University signals a potential paradigm shift, offering an early but crucial step toward overcoming this bottleneck. Led by Dr. Brian T. Chait’s Laboratory of Mass Spectrometry and Gaseous Ion Chemistry, the team has developed a novel prototype, dubbed MultiQ-IT, engineered to concurrently handle an immense number of molecules. This innovative design provides a foundational framework for constructing a new generation of mass spectrometry instruments capable of far greater speed and sensitivity, promising to revolutionize molecular analysis in much the same way parallel processing transformed fields like genomics and high-performance computing. The implications for scientific discovery, particularly in areas requiring comprehensive molecular profiling of highly complex biological systems, are profound, hinting at a future where the complete molecular tapestry of individual cells could be routinely mapped and thousands of chemical reactions monitored simultaneously.
The underlying principle of mass spectrometry involves ionizing molecules, imparting an electric charge, and then precisely measuring their mass-to-charge ratio. This fundamental measurement allows scientists to identify and quantify the specific substances present. While the technique itself is remarkably powerful, its traditional sequential operational mode presents a significant hurdle, especially when dealing with samples characterized by extreme molecular diversity and vast differences in abundance. Consider, for instance, the ambitious goals of single-cell proteomics and metabolomics, which aim to catalog every protein or metabolite within a single cell. Unlike DNA, these biomolecules cannot be amplified or copied, and their concentrations can vary by many orders of magnitude. The faint signals emanating from crucial, low-abundance molecules are frequently overwhelmed by the dominant signals from more prevalent species, akin to trying to hear a whisper in a roaring crowd. Current mass spectrometry sensitivity, while impressive, often falls short in these demanding scenarios, leading to incomplete or biased molecular snapshots.
For decades, this operational constraint has been a source of frustration for scientists like Dr. Chait, who recognized the latent, untapped capabilities of mass spectrometry. He envisioned a future where the technique could transcend its current limitations, performing at a scale previously unimaginable. The conceptual inspiration for this advancement came from the transformative power of "massive parallelization," a strategy that has already reshaped other scientific and technological domains. In the realm of computing, the development of Graphics Processing Units (GPUs) enabled the decomposition of complex problems into countless smaller tasks, processed concurrently, leading to exponential gains in performance essential for modern applications like artificial intelligence and scientific simulations. Similarly, the field of DNA sequencing underwent a dramatic revolution not by altering the fundamental chemistry of nucleotide identification, but by developing methodologies to execute millions of sequencing reactions in parallel. This shift drastically reduced the cost and time required to sequence an entire genome, moving from a billion-dollar endeavor to a routine analysis costing mere hundreds of dollars. The core idea, as Dr. Chait often emphasizes, was not to reinvent the underlying chemical reactions but to radically optimize the way they are performed.
The challenge, as articulated by senior research associate Andrew Krutchinsky, lay in translating this "obvious idea" of parallelization into a practical design for mass spectrometry. The solution eventually emerged from an unexpected source: the intricate biological machinery governing molecular transport within cells. Specifically, the researchers drew inspiration from nuclear pore complexes, which regulate the passage of molecules into and out of a cell’s nucleus. Rather than relying on a single, congested pathway, these complexes efficiently manage traffic across numerous small, distributed openings. The team posited whether mass spectrometry could be re-engineered to mimic this biological elegance, distributing the analytical load across multiple channels rather than funneling everything through a single point.
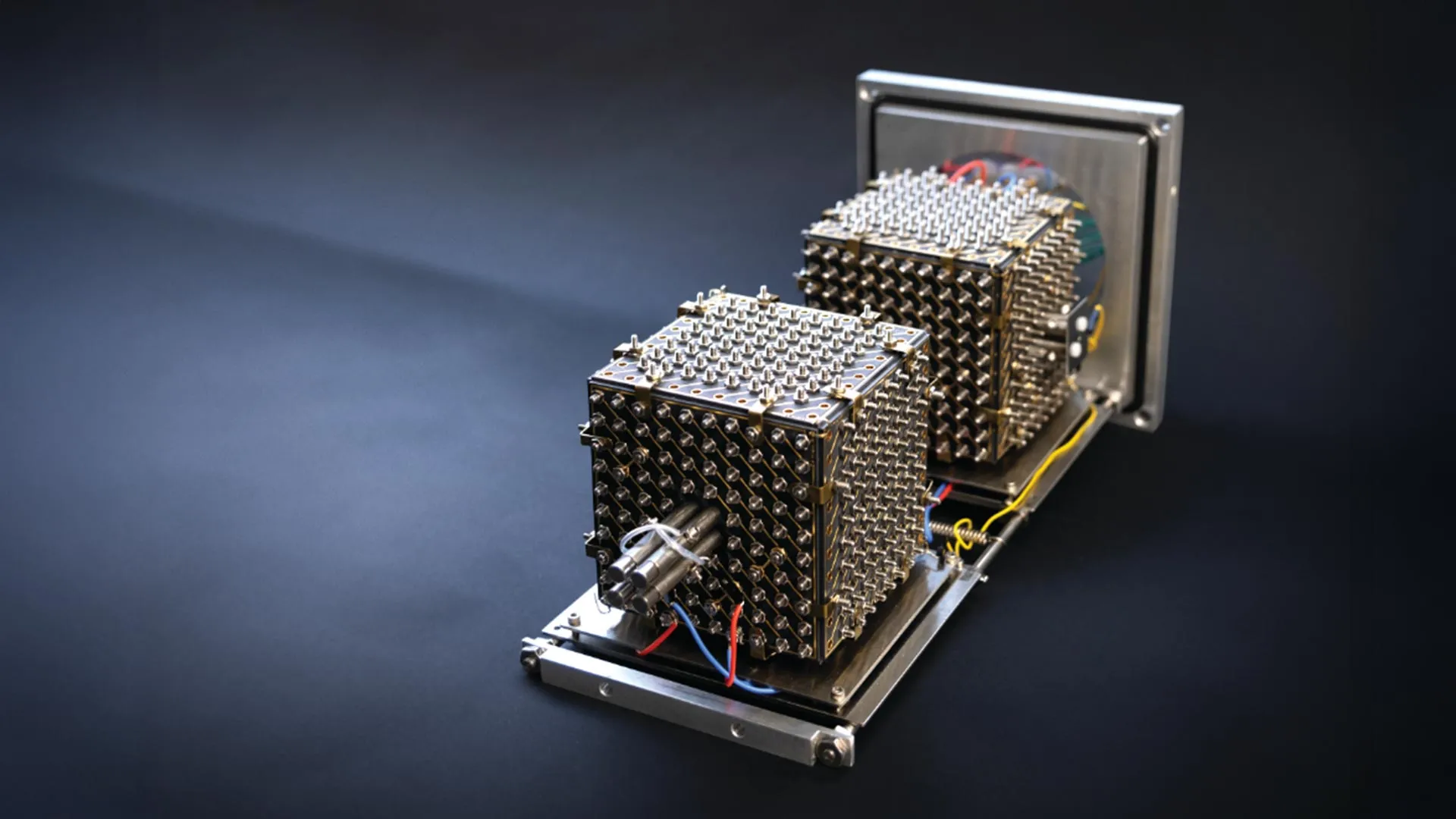
This conceptual leap culminated in the design of the MultiQ-IT’s core innovation: a sophisticated ion-trapping chamber intended to replace critical components found in conventional mass spectrometers. This unique, cube-shaped device is characterized by hundreds, and potentially thousands, of minute, electrically controlled apertures. Upon entering this chamber, ions undergo collisions with gas molecules, causing them to decelerate and move in a random fashion. This dynamic environment is precisely engineered to facilitate the simultaneous sorting, containment, and directed manipulation of multiple distinct groups of ions. Unlike traditional traps that process ions sequentially, the MultiQ-IT’s architecture allows for a multi-channel approach, vastly increasing throughput.
The development process involved iterative refinement, starting with a limited number of openings and progressively expanding the design. The team successfully scaled the system from just six apertures to over a thousand, meticulously testing its efficacy in managing and separating ion populations. This rigorous testing demonstrated the prototype’s ability to take a single incoming stream of ions and intelligently divide it into numerous parallel streams, each destined for simultaneous analysis. This parallel processing capability inherently addresses the "space charge effect," a critical limitation in conventional ion traps where the strong electrical repulsion among a high concentration of similarly charged particles restricts the maximum number of ions that can be held and analyzed effectively. By distributing ions across many channels, the MultiQ-IT significantly mitigates this repulsion, allowing for a far greater ion capacity without compromising analytical precision.
The performance metrics achieved by the MultiQ-IT prototype have been nothing short of impressive, underscoring its potential to redefine mass spectrometry capabilities. A version equipped with 486 ports demonstrated the capacity to hold up to ten billion charges concurrently—a staggering thousand-fold increase compared to the typical capacity of conventional ion traps. Beyond sheer capacity, the system introduces a groundbreaking approach to enhancing detection sensitivity. It intelligently leverages a small electrical voltage barrier positioned at the exits of the trap. This barrier is calibrated to allow singly charged ions, which often represent less informative background noise, to escape the system. Conversely, multiply charged ions, frequently indicative of more complex and biologically significant molecules, are retained within the trap for analysis. This selective expulsion of background interference dramatically elevates the signal-to-noise ratio, achieving improvements of up to 100-fold. Such a substantial boost in sensitivity opens the door to detecting proteins and other biomolecules that were previously obscured by noise, rendering them effectively undetectable.
Further experiments with a larger iteration of the design, featuring 1,134 ports, revealed remarkable efficiency in its filtering capabilities. Researchers observed that only a modest 39 open ports were required to achieve half of the system’s maximum filtering efficiency. This observation strikingly mirrors the operational efficiency of biological systems, such as cells utilizing a limited number of nuclear pores to manage vast molecular traffic. This increased sensitivity has direct implications for critical research areas, including the enhanced detection of low-abundance crosslinked peptides. These specific peptides are invaluable for meticulously mapping the three-dimensional structures of large protein complexes, providing crucial insights into their function and interactions. As Krutchinsky points out, "The least abundant things can be more important than the more abundant things," underscoring the significance of a technology that can bring these elusive molecules into focus.
At its current stage, the MultiQ-IT remains a proof of concept, a foundational design demonstrating what is technologically achievable rather than a finalized commercial product. However, its significance lies in laying down a clear blueprint for the next generation of mass spectrometry instruments. The journey from a groundbreaking scientific discovery to a widely adopted technological tool is often extensive, spanning years or even decades. The first transistor, for example, heralded a revolution, yet it took many years to transition to integrated circuits containing billions of transistors on a single chip. Similarly, the initial discovery of DNA sequencing reactions preceded modern genomics by decades. In both these transformative cases, the crucial first step was demonstrating that the envisioned capability was, indeed, possible.
The researchers at Rockefeller University believe they have now provided that pivotal demonstration for mass spectrometry, showcasing a viable pathway toward dramatically more efficient and powerful molecular analysis. This development holds immense promise for accelerating discovery in numerous fields. In drug development, it could enable faster screening of potential therapeutic compounds and a more detailed understanding of drug-target interactions. For biomarker identification, it could lead to earlier and more accurate disease diagnostics, paving the way for truly personalized medicine. In fundamental biological research, the ability to profile cellular contents with unprecedented depth and breadth will unlock deeper insights into disease mechanisms, cellular signaling, and metabolic pathways. The MultiQ-IT represents not just an incremental improvement, but a foundational innovation that could usher in a new era of molecular science, pushing the boundaries of what we can see, understand, and ultimately, achieve in the complex world of molecules.