The intricate machinery of human cells relies on a meticulously orchestrated set of genetic instructions that govern their growth, division, and specialized functions. When this delicate genetic blueprint suffers disruptions, particularly through the accumulation of errors in its chromosomal architecture, the groundwork for diseases like cancer can be laid. These genetic inaccuracies, known as chromosomal abnormalities, represent critical early indicators of cellular dysfunction, propelling otherwise healthy cells down a perilous path toward malignancy. Such defects, encompassing alterations in the number or structural integrity of chromosomes, are increasingly recognized as fundamental drivers of aggressive forms of cancer, strongly correlated with patient mortality, metastatic spread, disease recurrence, and resistance to chemotherapy treatments.
For over a hundred years, the scientific community has pondered the profound connection between these abnormal chromosomes and the genesis of cancer. It was in the nascent years of the twentieth century that German biologist Theodor Boveri, meticulously observing dividing cells under the microscope, first posited this revolutionary idea. His pioneering work, published in 1914, led him to hypothesize that an improper chromosomal complement within a cell could be the root cause of its uncontrolled proliferation, thus proposing the "Somatic Mutation Theory" of cancer. Boveri’s insights were remarkably prescient, especially considering the rudimentary state of genetic understanding at the time; the very structure of DNA would not be elucidated for another half-century. Despite the compelling nature of his theory, direct experimental validation remained elusive for generations of researchers due to significant technical limitations.
The primary hurdle in substantiating Boveri’s hypothesis lay in the inherent difficulty of studying chromosomal abnormalities. These critical cellular errors are transient and relatively rare events; at any given moment, only a minuscule fraction of cells within a population might display such defects. Furthermore, biological systems possess robust quality control mechanisms, meaning many cells with significant chromosomal errors are naturally eliminated through programmed cell death (apoptosis) or immune surveillance. Historically, scientists attempting to investigate these elusive events were forced to rely on painstaking manual microscopy, a labor-intensive and incredibly slow process that permitted the isolation and study of only a handful of cells at a time. This methodological bottleneck severely constrained the scope and pace of research, making it nearly impossible to gather the statistical evidence needed to fully understand the frequency and conditions under which these abnormalities arise.
A significant leap forward in addressing this long-standing challenge has now been achieved by researchers within the Korbel Group at the European Molecular Biology Laboratory (EMBL) Heidelberg. They have engineered a sophisticated artificial intelligence (AI)-driven platform, which they named machine learning-assisted genomics and imaging convergence (MAGIC). This groundbreaking system represents a powerful fusion of advanced microscopy, single-cell genomic analysis, and cutting-edge AI algorithms, designed to overcome the historical limitations in studying the origins of chromosomal errors. By precisely identifying the cellular conditions that foster these genetic mishaps, MAGIC promises to fundamentally reshape our understanding of how cancer takes hold at its earliest stages.
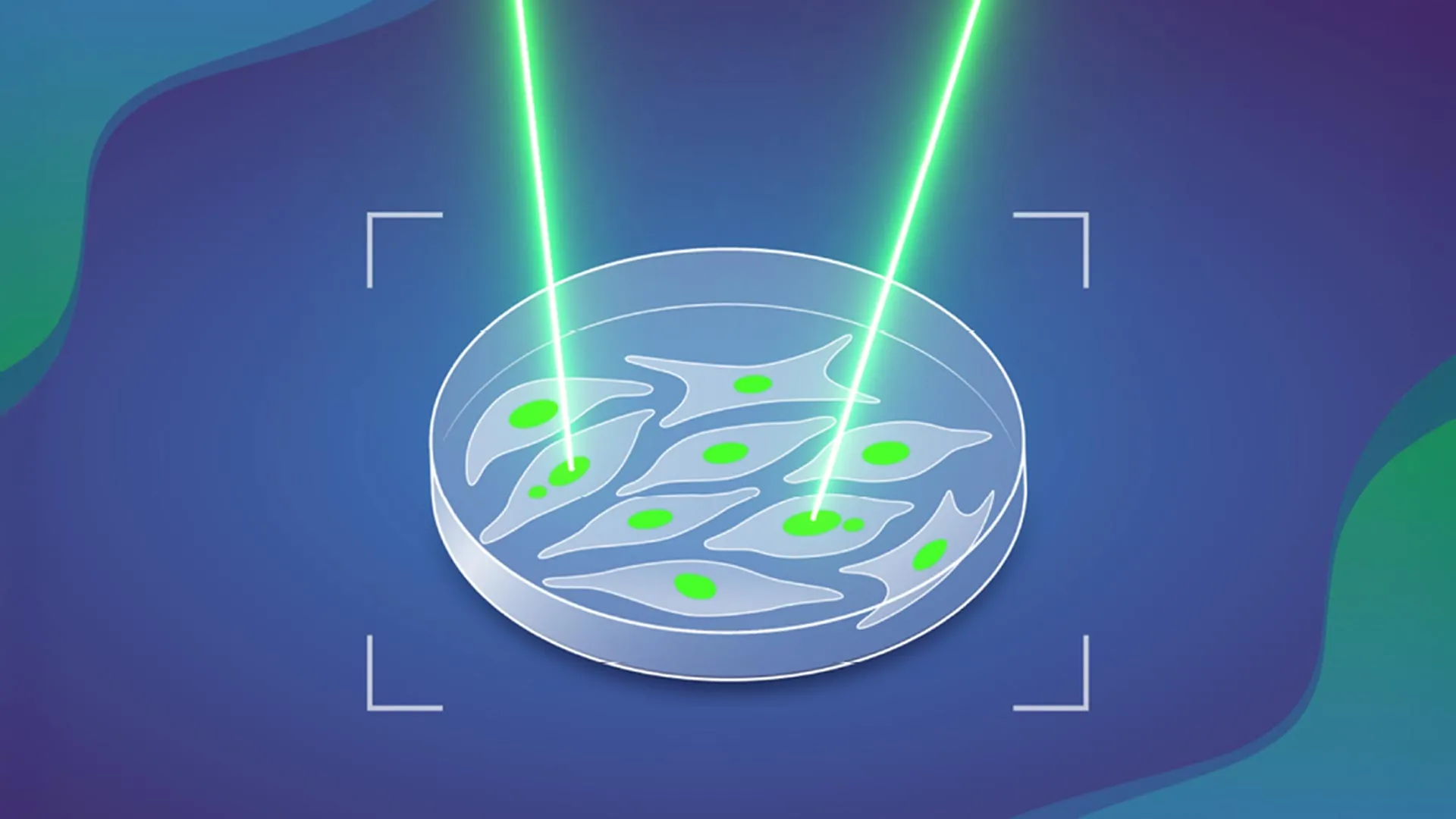
The genesis of the MAGIC system stemmed from a collaborative effort led by Marco Cosenza, a Research Scientist in the Korbel Group. Recognizing the shared technical obstacles faced by various EMBL teams in their cellular investigations, Cosenza embarked on designing an automated solution. His vision was to create a platform that could mimic the selective power of human observation but at an unprecedented scale and speed. The eventual system operates with a functional elegance akin to an automated "laser tag" game for cells, where specific cellular features trigger a targeted response.
Central to MAGIC’s operation is its ability to detect ‘micronuclei’ – small, distinct compartments within a cell that contain fragmented pieces of DNA separate from the main nucleus. The presence of micronuclei is a widely recognized biomarker indicating genomic instability and a heightened propensity for cells to develop further chromosomal aberrations, thereby increasing their likelihood of malignant transformation. When MAGIC’s integrated AI identifies a cell containing one or more micronuclei, it precisely targets and marks that cell using a sophisticated laser mechanism. This marking process leverages a photoconvertible dye, a remarkable fluorescent molecule that undergoes a permanent change in its light emission spectrum upon exposure to a specific wavelength of light. This irreversible "tag" ensures that the identified cells can be distinguished and isolated for subsequent in-depth analysis.
The operational workflow of the MAGIC system unfolds through a series of seamlessly integrated, automated steps. Initially, an automated microscope systematically captures a vast array of high-resolution images from a cellular sample. These images are then fed into a highly specialized machine learning algorithm. This algorithm has been rigorously trained using extensive datasets of manually annotated images, where human experts meticulously labeled cells exhibiting micronuclei. This supervised learning approach enables the AI to accurately discern and identify these subtle yet critical cellular features with high fidelity.
Upon detecting a micronucleus-containing cell, the algorithm relays its precise coordinates back to the automated microscope. The microscope then precisely directs a focused beam of light at that specific cell, activating the photoconvertible dye and permanently tagging it. Once tagged, these living cells can be efficiently separated from the untagged population using advanced cell sorting techniques such as flow cytometry, which rapidly analyzes and isolates cells based on their fluorescent properties. The isolated, "pre-cancerous" cells are then subjected to detailed molecular investigation, including single-cell genomic sequencing, allowing researchers to scrutinize their complete genetic makeup and pinpoint the exact nature of their chromosomal abnormalities. This high-throughput approach dramatically accelerates research, enabling the analysis of close to 100,000 cells in under a day, a scale previously unimaginable through manual methods.
Applying the MAGIC system to cultured cells derived from normal human tissues yielded profound insights into the kinetics of chromosomal abnormalities. The research team discovered that, under normal physiological conditions, approximately 10% of all cell divisions spontaneously result in some form of chromosomal error. This baseline rate provides a crucial quantitative measure of inherent genomic instability. Even more significantly, their investigations revealed a stark acceleration in this rate when the gene encoding p53, a renowned tumor suppressor protein often referred to as the "guardian of the genome," was mutated. In cells with a compromised p53 pathway, the frequency of spontaneous chromosomal abnormalities nearly doubled, underscoring the critical role of p53 in maintaining genomic integrity and preventing the accumulation of cancer-driving mutations. The team also explored other influencing factors, such as the presence and precise location of double-stranded DNA breaks within chromosomes, further illuminating the complex interplay of factors contributing to genomic instability.
The implications of the MAGIC platform extend far beyond the immediate findings on cancer initiation. Its inherent flexibility and adaptability make it a powerful tool with broad potential across numerous biological disciplines. While the initial study focused on training the AI to recognize micronuclei, the underlying machine learning architecture can be readily adapted to identify a wide array of other visually distinguishable cellular features. As Jan Korbel, senior scientist at EMBL and senior author of the paper published in Nature, articulated, "As long as you have a feature that can be discriminated visually from a ‘regular’ cell, you can — thanks to AI — train the system to detect it." This versatility opens avenues for breakthroughs in diverse fields, including drug discovery, developmental biology, regenerative medicine, and the study of aging, by enabling high-throughput identification and isolation of cells exhibiting specific phenotypes.
This ambitious research project was a testament to the power of interdisciplinary collaboration, drawing expertise from various institutions and groups. Key contributions came from within EMBL, including the Advanced Light Microscopy Facility (ALMF) and the Pepperkok Team at EMBL Heidelberg, as well as Isidro Cortes-Ciriano’s group at EMBL-EBI. External collaborators included Andreas Kulozik’s team at the German Cancer Research Centre (DKFZ), which is also a partner in the Molecular Medicine Partnership Unit (MMPU) between EMBL and the University of Heidelberg. Such synergistic efforts highlight the collaborative spirit essential for tackling complex biological questions in contemporary science. The advent of the MAGIC system not only provides compelling empirical validation for a century-old theory about cancer’s origins but also establishes a transformative technological framework poised to accelerate discoveries across the vast landscape of biological research, ultimately bringing us closer to understanding, preventing, and treating devastating diseases.