A groundbreaking scientific endeavor, involving collaborative efforts from the Salk Institute for Biological Studies, the UNC Lineberger Comprehensive Cancer Center, and UC San Diego, has illuminated fundamental genetic mechanisms that dictate the fate of critical immune sentinels. These specialized lymphocytes, designated as CD8 "killer" T cells, possess the remarkable capacity to either evolve into persistent defenders offering enduring protection against pathogens and malignancies or to succumb to a debilitating state of functional decline known as exhaustion. The pivotal discovery emanating from this research posits that the inactivation of a mere two specific genes can effectively reawaken the dormant anti-tumor capabilities of these fatigued immune cells, thereby restoring their ability to mount a vigorous assault on cancerous growths.
This seminal work, meticulously detailed in the esteemed scientific journal Nature, establishes a foundational blueprint with the potential to empower scientists in the deliberate engineering of T cells. The objective is to cultivate immune cells that can concurrently retain robust long-term immunological memory and sustain potent cancer-eliminating activity. The ramifications of these findings extend far beyond the realm of oncology, promising significant advancements in the therapeutic landscape for a spectrum of infectious diseases characterized by persistent viral burdens or chronic inflammation.
At the forefront of cellular defense, CD8 killer T cells play an indispensable role in orchestrating the immune system’s response by actively identifying and eradicating cells compromised by viral invasion or cancerous transformation. However, in the face of protracted infections or persistent tumor burdens, these vigilant cells can gradually experience a diminution in their efficacy. Over an extended period, this attrition can precipitate a state of profound dysfunction, commonly referred to as T cell exhaustion, wherein their capacity to neutralize threats suffers a considerable decline.
Constructing a Comprehensive Genetic Cartography of T Cell Phenotypes
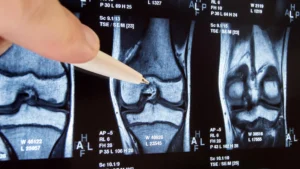
The inherent challenge in distinguishing between protective T cells and their exhausted counterparts lies in their superficial resemblance; under conventional analytical methods, they often appear virtually indistinguishable. To surmount this diagnostic hurdle, the research team embarked on an exploration to ascertain whether divergent cellular states could be demarcated based on their underlying genetic activity.
A significant leap forward was achieved through the meticulous construction of an intricate genetic atlas. This comprehensive map chronicles a wide array of CD8 T cell states, illustrating the dynamic spectrum along which these immune cells transition, ranging from a highly protective and functional phenotype to one that is severely impaired and dysfunctional.
"Our overarching aspiration is to elevate the efficacy of immune-based therapies by formulating precise ‘recipes’ for the optimal design of T cells," articulated co-corresponding author Susan Kaech, PhD, who held a professorship at the Salk Institute during the study’s progression. "To achieve this ambitious goal, our initial imperative was to meticulously identify the specific molecular components that exhibit unique activity within one T cell state but remain quiescent in others. By developing an exhaustive atlas of CD8 T cell states, we successfully pinpointed the critical factors that define programs associated with protection versus those leading to dysfunction—information that is absolutely essential for the precise engineering of robust and effective immune responses."
The Reversibility of T Cell Exhaustion: A Paradigm Shift
To unravel the intricate regulatory pathways governing these distinct immune states, the researchers subjected nine discrete CD8 T cell conditions to rigorous examination. This comprehensive investigation employed sophisticated laboratory techniques, advanced genetic manipulation tools, meticulously controlled animal models, and extensive computational analysis. Their endeavors yielded the identification of several transcription factors—proteins that act as molecular switches to modulate gene expression—which serve as crucial determinants, guiding T cells toward either sustained functionality or the state of exhaustion.
Among the identified regulators, the scientists pinpointed two transcription factors, ZSCAN20 and JDP2, which had not previously been implicated in the pathogenesis of T cell exhaustion. Crucially, when the genes encoding these factors were experimentally disabled, the exhausted T cells exhibited a remarkable recovery of their tumor-killing prowess, while simultaneously preserving their capacity for long-term immune memory.
"We deliberately manipulated specific genetic switches within the T cells to assess whether we could reinstate their tumor-eliminating function without compromising their ability to confer long-lasting immunological protection," explained co-corresponding author H. Kay Chung, PhD, an assistant professor at UNC Lineberger, who initiated this research at the Salk Institute prior to her tenure at UNC. "Our observations confirmed that it is indeed feasible to decouple these two critical outcomes, separating the capacity for robust effector function from the maintenance of long-term memory."
These findings represent a significant challenge to a long-held tenet in immunology, which posited that immune exhaustion is an inevitable and irreversible consequence of prolonged and intense immune system activation.
Engineering Superior Immune Cells for Advanced Cancer Therapeutics
The researchers posit that the genetic atlas they have meticulously constructed serves as an invaluable guide for the rational design of more potent immune cells, particularly for application in advanced therapeutic modalities such as adoptive cell transfer (ACT) and chimeric antigen receptor (CAR) T cell therapy.
"With this detailed map in hand, we can now provide T cells with far more explicit instructions, thereby assisting them in retaining the traits that enable them to effectively combat cancer or infection over extended durations, while simultaneously avoiding the molecular pathways that precipitate their burnout," stated Dr. Kaech. "By successfully disentangling these two distinct cellular programs, we are on the cusp of being able to design immune cells that are both exceptionally durable and highly effective in the context of cancer treatment and chronic infectious diseases."
This groundbreaking discovery holds particular significance for the treatment of solid tumors, a clinical challenge where the pervasive nature of immune exhaustion frequently serves as a formidable impediment to therapeutic success.
The Synergy of Artificial Intelligence and Precision Immune Engineering
Looking ahead, the research team intends to integrate cutting-edge experimental methodologies with AI-driven computational modeling. Their ambitious objective is to develop a considerably expanded repertoire of highly precise genetic "recipes" designed to program T cells into specific functional states, thereby enhancing the precision and efficacy of cellular therapies.
"Given that genes operate within intricate regulatory networks that are notoriously difficult to decipher, powerful computational tools are indispensable for accurately identifying which specific regulators are instrumental in driving particular cell states," observed co-corresponding author Wei Wang, PhD, a distinguished professor at UC San Diego. "This study unequivocally demonstrates that we are now capable of precisely manipulating the fates of immune cells, thereby unlocking novel avenues for the enhancement of immunotherapies."
By elucidating the intricate decision-making processes by which killer T cells navigate the dichotomy between resilience and exhaustion, this research brings scientists considerably closer to the ability to deliberately steer immune responses, rather than passively witnessing their deterioration in the face of prolonged disease. The collaborative effort involved numerous other researchers including Eduardo Casillas, Ming Sun, Shixin Ma, Shirong Tan, Brent Chick, Victoria Tripple, Bryan McDonald, Qiyuan Yang, Timothy Chen, Siva Karthik Varanasi, Michael LaPorte, Thomas H. Mann, Dan Chen, Filipe Hoffmann, Josephine Ho, April Williams, and Diana C. Hargreaves from Salk; Cong Liu, Alexander N. Jambor, Z. Audrey Wang, Jun Wang, Zhen Wang, Jieyuan Liu, and Zhiting Hu from UC San Diego; Anamika Battu, Brandon M. Pratt, Fucong Xie, Brian P. Riesenberg, Elisa Landoni, Yanpei Li, Qidang Ye, Daniel Joo, Jarred Green, Zaid Syed, Nolan J. Brown, Matthew Smith, Jennifer Modliszewski, Yusha Liu, Ukrae H. Cho, Gianpietro Dotti, Barbara Savoldo, Jessica E. Thaxton, and J. Justin Milner from UNC; Peixiang He, Longwei Liu, and Yingxiao Wang from the University of Southern California; and Yiming Gao from Texas A&M University. This extensive research initiative was generously supported by funding from the National Institutes of Health (grants R37AI066232, R01AI123864, R21AI151986, R01CA240909, R01AI150282, R01HG009626, K01EB034321, R01AI177864, R01CA248359, R01CA244361, AI151123, EB029122, GM140929) and the Damon Runyon Cancer Research Foundation.