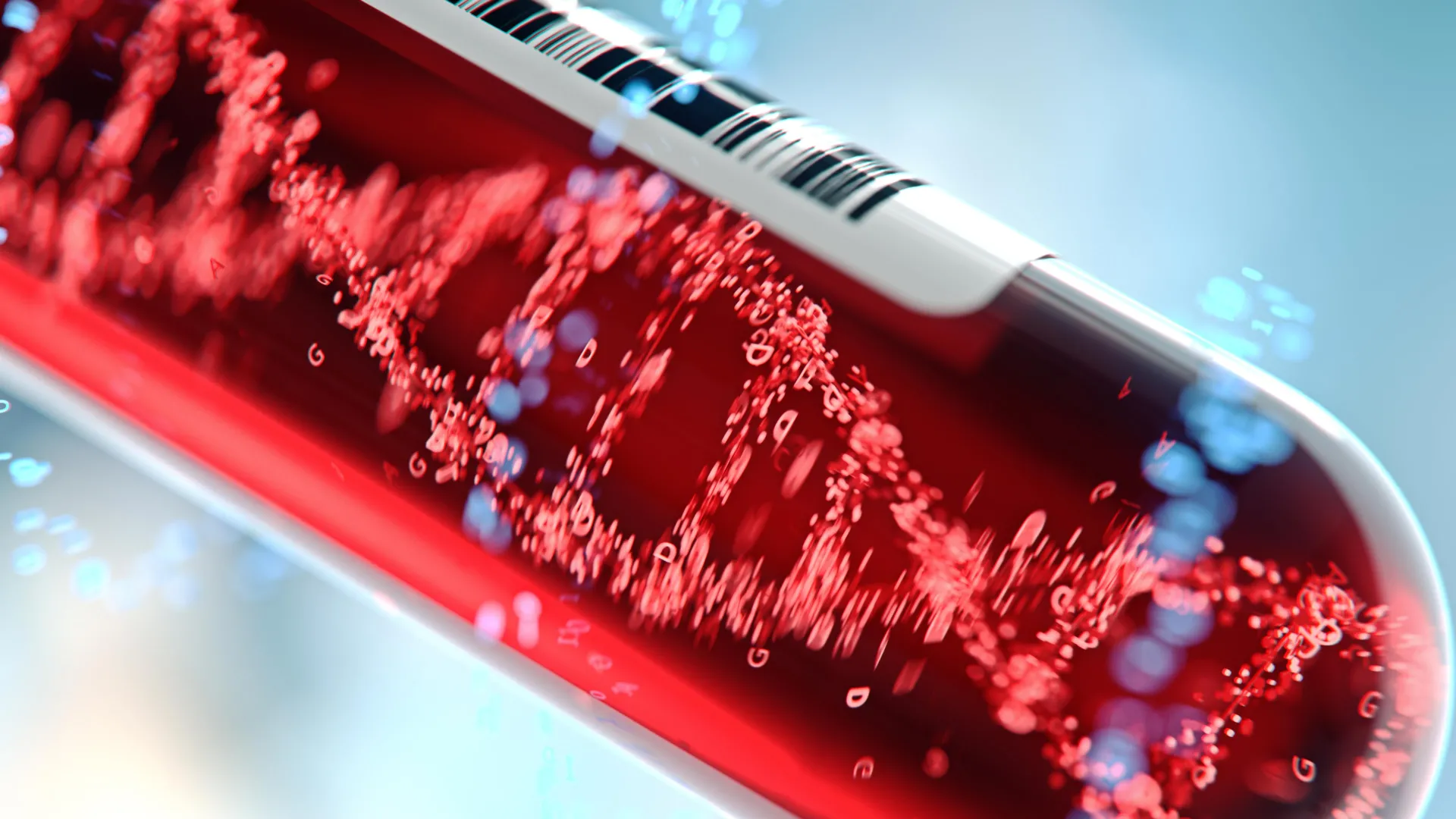
A groundbreaking advance in medical diagnostics, spearheaded by researchers at the Johns Hopkins Kimmel Cancer Center, has unveiled a sophisticated artificial intelligence-driven liquid biopsy capable of identifying the subtle molecular signatures of liver fibrosis and cirrhosis years before clinical symptoms manifest. This innovative approach, detailed in a recent publication in Science Translational Medicine, marks a significant expansion of liquid biopsy applications beyond their traditional focus on cancer detection, opening new avenues for the early diagnosis and intervention in a range of chronic illnesses. By meticulously analyzing the genome-wide patterns of cell-free DNA (cfDNA) fragments found in the bloodstream, this technology offers a non-invasive window into a person’s physiological state, potentially revolutionizing how silent diseases are identified and managed.
For decades, the diagnosis of chronic liver conditions like fibrosis and cirrhosis has presented considerable challenges. Liver fibrosis, the initial stage of scarring, is often asymptomatic, meaning individuals experience no outward signs or discomfort. This "silent" progression frequently allows the disease to advance undetected, ultimately leading to cirrhosis—a more severe, irreversible form of scarring that impairs liver function and significantly increases the risk of liver cancer, also known as hepatocellular carcinoma. Current diagnostic methods for liver disease are often invasive, such as liver biopsies, which carry risks and are not practical for widespread screening. Non-invasive options, including traditional blood tests for liver enzymes or imaging techniques like specialized ultrasound and magnetic resonance elastography, frequently lack the sensitivity to detect early-stage fibrosis or may not be readily accessible to all patients. The limitations of these existing tools mean that millions of individuals at risk remain undiagnosed until their condition has progressed to a point where treatment options are more limited.
The core of this new diagnostic paradigm lies in "fragmentome technology," a method that examines the intricate ways DNA fragments are shed into the bloodstream and their subsequent patterns across the entire human genome. When cells in the body die, they release fragments of their DNA into the circulating blood plasma. While the presence of cfDNA has been known for some time, this research moves beyond simply detecting its existence or looking for specific mutations. Instead, it delves into the nuanced characteristics of these fragments: their size, their precise locations within the genome, and their overall distribution. These patterns, researchers hypothesize, act as distinct fingerprints, offering clues about the health and activity of various tissues and organs, including the liver.
In a comprehensive study, scientists performed whole genome sequencing on cfDNA samples collected from 1,576 individuals. This diverse cohort included patients diagnosed with various stages of liver disease, as well as those with other medical conditions. Unlike many liquid biopsy platforms that target specific genetic alterations or epigenetic modifications often associated with cancer, the fragmentome approach casts a much wider net. It scrutinizes approximately 40 million cfDNA fragments per sample, spanning thousands of genomic regions, including those repetitive DNA sequences that have historically been under-explored. This generates an immense dataset, far larger and more complex than what typical liquid biopsies analyze.
To make sense of this deluge of information, the research team employed sophisticated machine learning algorithms. These algorithms were trained to identify specific fragmentation patterns that correlated with different stages of liver disease. The result was the development of a highly sensitive classification system capable of distinguishing early liver disease, advanced fibrosis, and full-blown cirrhosis. Dr. Victor Velculescu, co-director of the cancer genetics and epigenetics program at the Johns Hopkins Kimmel Cancer Center and a co-senior author of the study, emphasized the foundational nature of this work. He noted that this builds directly on earlier fragmentome research in oncology, but with a novel application to chronic non-cancerous diseases, highlighting liver fibrosis and cirrhosis as critical examples where early intervention can profoundly alter disease trajectory.
The clinical implications of such an early detection system are profound. Liver fibrosis, in its nascent stages, is often reversible with appropriate lifestyle changes or medical treatments for underlying causes (such as fatty liver disease or chronic viral hepatitis). However, if left unchecked, it inexorably progresses to irreversible cirrhosis and an elevated risk of liver cancer. With an estimated 100 million Americans living with liver conditions that predispose them to cirrhosis and liver cancer, the ability to identify these conditions proactively could prevent countless cases of advanced disease and associated mortality. Dr. Velculescu underscored this point, stating that detecting precursor conditions early could enable physicians to address underlying pathologies sooner, potentially halting progression and averting cancer development.
A key differentiator of this fragmentome analysis, as explained by Dr. Akshaya Annapragada, the study’s first author and an M.D./Ph.D. student in the Velculescu lab, is its holistic perspective. Rather than searching for isolated genetic mutations, which might be present only in certain cancers, the method analyzes the entire fragmentome, providing a comprehensive snapshot of a person’s physiological state. This expansive data, combined with advanced machine learning, empowers the creation of highly specific classifiers for a multitude of health conditions, each distinct and non-cross-reactive. For instance, a classifier for liver fibrosis would be uniquely identifiable from a cancer classifier, even though both are derived from the same underlying platform. This specificity is crucial for accurate diagnosis and avoids confounding signals between different disease states.
The origins of this breakthrough can be traced back to a 2023 Cancer Discovery study, also led by Dr. Velculescu, which investigated the fragmentome in the context of liver cancer. During that research, scientists observed subtle, yet distinct, DNA signals in patients with fibrosis or cirrhosis, even when their overall fragmentation profiles appeared largely normal. This intriguing observation prompted the dedicated investigation into fragmentome patterns specifically associated with non-cancerous liver fibrosis and cirrhosis, ultimately leading to the current findings.
Beyond liver disease, the research also explored the broader utility of fragmentome technology. In a separate analysis involving 570 individuals suspected of having serious illnesses, the team developed a "fragmentation comorbidity index." This index effectively differentiated between individuals with high and low Charlson Comorbidity Index scores—a widely recognized clinical metric used to estimate how co-existing health conditions impact a patient’s mortality risk. Remarkably, the fragmentome-based index independently predicted overall survival and, in some instances, demonstrated greater specificity than conventional inflammatory markers. Certain fragmentation signatures were also found to correlate with poorer clinical outcomes, suggesting the fragmentome could serve as a powerful prognostic tool, offering insights into a patient’s overall health and resilience.
While the current study primarily focused on liver disease, the researchers observed preliminary fragmentome signals associated with other significant medical conditions, including cardiovascular, inflammatory, and neurodegenerative disorders. Although the study cohort was not sufficiently large to develop distinct disease classifiers for each of these conditions, these findings strongly suggest the technology’s potential for broader applicability across various chronic illnesses. This paves the way for future research to investigate and validate fragmentome signatures for a wider spectrum of diseases, potentially leading to a new era of proactive and preventative healthcare.
It is important to note that the liver fibrosis assay described in this study remains a prototype. Before it can be introduced as a clinical test, further rigorous refinement and extensive validation are required. The next phases of this research will involve validating the liver disease classifier in larger and more diverse patient populations, as well as systematically exploring fragmentome signatures linked to other chronic conditions. This meticulous process ensures the test’s reliability, accuracy, and clinical utility across different demographics and healthcare settings.
The substantial research effort behind this innovation was supported by a consortium of funding bodies, including the National Institutes of Health, the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation, the SU2C in-Time Lung Cancer Interception Dream Team Grant, and the Gray Foundation, among others. This multi-institutional and multi-funder collaboration underscores the significant scientific and societal value placed on developing advanced, non-invasive diagnostic tools for chronic diseases. The pioneering work from the Johns Hopkins Kimmel Cancer Center team represents a pivotal step towards a future where early detection of "silent" chronic diseases becomes routine, enabling timely interventions that can profoundly improve patient outcomes and alleviate the burden on healthcare systems worldwide.