A groundbreaking initiative by researchers at Rice University has culminated in the creation of the inaugural comprehensive, label-free molecular atlas of an Alzheimer’s-affected brain within an animal model, offering an unprecedentedly granular perspective on the disease’s incipient stages and propagation mechanisms. The profound societal burden of Alzheimer’s, claiming more lives annually than breast and prostate cancers combined, amplifies the critical imperative to unravel the intricate biological drivers underpinning its relentless progression. This pioneering work, detailed in the esteemed journal ACS Applied Materials and Interfaces, transcends the conventional focus on amyloid plaques, illuminating a complex and heterogeneous landscape of chemical alterations distributed unevenly throughout the brain.
The scientific endeavor leveraged a sophisticated, light-based imaging modality coupled with advanced artificial intelligence algorithms to meticulously scrutinize brain tissue samples obtained from both healthy control subjects and those exhibiting Alzheimer’s pathology. This advanced analytical framework enabled the identification of subtle, yet significant, chemical deviations that are not exclusively localized to the hallmark amyloid deposits. Instead, these alterations manifest as intricate, non-uniform patterns, suggesting a more pervasive and multifaceted disruption of neuronal and glial function.
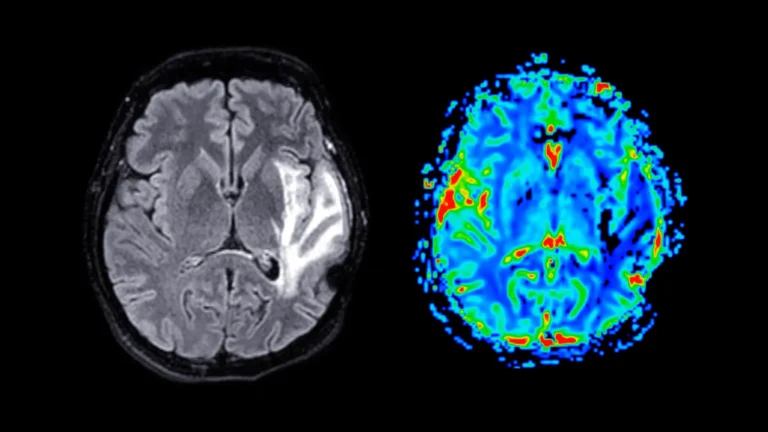
At the heart of this investigation lies hyperspectral Raman imaging, a cutting-edge spectroscopic technique that harnesses the power of a laser to discern the unique molecular signatures inherent to biological tissues. Unlike conventional Raman spectroscopy, which captures a singular data point of chemical information at a specific molecular locus, hyperspectral Raman imaging systematically acquires thousands of such measurements across an entire tissue section. This exhaustive data acquisition process constructs a comprehensive map, vividly illustrating the spatial variations in chemical composition across diverse brain regions. The researchers meticulously scanned entire brain specimens, slice by slice, amassing a vast repository of overlapping spectral data points. This meticulous approach facilitated the generation of high-resolution molecular maps, providing an unadulterated chemical portrait of both healthy and diseased neural tissue.
A critical advantage of this methodology is its "label-free" nature, obviating the need for potentially disruptive staining or labeling agents such as dyes, fluorescent proteins, or molecular tags. This inherent characteristic ensures that the observed molecular landscape accurately reflects the brain’s native chemical state, free from the confounding influences of exogenous compounds. As articulated by Ziyang Wang, a doctoral candidate in electrical and computer engineering at Rice and a lead author of the study, this unbiased approach is particularly well-suited for the discovery of novel disease-associated changes that might otherwise elude detection through conventional methods. The resulting data, an immense reservoir of chemical information, was subsequently subjected to rigorous analysis by sophisticated machine learning (ML) algorithms.
The integration of unsupervised ML techniques allowed the algorithms to autonomously identify emergent patterns within the spectral data without any preconceived notions or prior assumptions about the underlying biological processes. This initial phase of analysis facilitated the classification of tissue samples based purely on their intrinsic molecular characteristics. Subsequently, the researchers employed supervised ML, a process involving the training of models on labeled datasets to accurately differentiate between Alzheimer’s-affected and healthy brain tissue. This supervised approach proved instrumental in quantifying the degree to which specific brain regions exhibited Alzheimer’s-related chemical signatures, thereby highlighting areas of heightened vulnerability.
The insights gleaned from this machine learning analysis revealed a profound heterogeneity in the distribution of Alzheimer’s-induced changes across the brain. Certain brain regions demonstrated marked chemical alterations, indicative of significant pathological insult, while others remained comparatively less affected. This uneven distribution, the researchers posit, may offer a compelling explanation for the insidious, gradual onset of Alzheimer’s symptoms and could also shed light on the limited efficacy of therapeutic strategies that target a single pathological pathway.
Beyond the well-established accumulation of misfolded proteins, the study uncovered broader metabolic dysregulations that distinguish Alzheimer’s brains from their healthy counterparts. Significant regional variations in the concentrations of cholesterol and glycogen were observed, with the most pronounced discrepancies manifesting in brain areas critically involved in memory processing, notably the hippocampus and the cerebral cortex. Cholesterol plays a vital role in maintaining the structural integrity and functional fluidity of neuronal membranes, while glycogen serves as a readily accessible intracellular energy reserve. The co-occurrence of altered cholesterol and glycogen levels in memory-centric regions suggests that Alzheimer’s pathology extends beyond protein aggregation to encompass fundamental disruptions in brain cell architecture and energy homeostasis, as explained by Shengxi Huang, an associate professor of electrical and computer engineering and materials science and nanoengineering, and the study’s corresponding author. Huang, also affiliated with several prominent research institutes at Rice, emphasized that these findings collectively support a more holistic understanding of Alzheimer’s, one that acknowledges pervasive disturbances in brain structure and energy metabolism in addition to proteinopathy.
The genesis of this ambitious project emerged from ongoing dialogues concerning novel methodologies for investigating the intricate complexities of the Alzheimer’s brain. Initially, the research focused on analyzing comparatively smaller tissue areas. However, a pivotal conceptual shift occurred when the team considered the potential of mapping the entire brain to achieve a more expansive and holistic understanding of the disease’s molecular footprint. The successful integration of advanced measurement techniques and sophisticated analytical algorithms was the culmination of numerous iterative cycles of experimentation and refinement.
Upon the successful compilation of the complete chemical map, the implications were immediately apparent. Previously imperceptible patterns began to emerge, offering a level of detail that had been unattainable with conventional imaging approaches. The realization of these findings, revealing a hidden stratum of molecular information that had long awaited the development of appropriate analytical tools, was profoundly rewarding for the research team.
By providing the first detailed, dye-free chemical maps of the Alzheimer’s brain, this research offers a paradigm shift in our comprehension of the disease. The scientists express optimism that these findings will pave the way for earlier and more accurate diagnostic methods and will ultimately contribute to the development of more efficacious strategies aimed at slowing disease progression. The research was generously supported by grants from the National Science Foundation, the National Institutes of Health, and the Welch Foundation, underscoring the collaborative and well-funded nature of this significant scientific undertaking.