A significant advancement in understanding the intricate molecular underpinnings of Alzheimer’s disease has been achieved by a research consortium led by investigators at the University of California, Irvine’s Joe C. Wen School of Population & Public Health. This pioneering work has culminated in the creation of the most detailed topographical maps to date, illustrating the direct influence one gene exerts upon another within the specific cellular environment of the Alzheimer’s-affected brain. These sophisticated maps transcend the mere identification of correlated gene activity, instead delving into the active regulatory hierarchies that dictate gene expression across diverse brain cell populations.
The cornerstone of this breakthrough is a novel machine learning framework developed by the research team, christened SIGNET. Traditional bioinformatics tools often rely on identifying genes whose expression patterns fluctuate in unison, inferring a connection based on co-occurrence. SIGNET, however, is engineered with a fundamentally different objective: to discern genuine causal linkages, pinpointing which genes are not merely associated but are actively orchestrating the behavior of others. Employing this advanced methodology, the researchers have illuminated crucial biological pathways implicated in the progressive cognitive decline and neuronal degradation characteristic of Alzheimer’s disease.
The groundbreaking findings of this study have been formally disseminated in the peer-reviewed publication, Alzheimer’s & Dementia: The Journal of the Alzheimer’s Association. Beyond elucidating fundamental disease mechanisms, the research also introduces a roster of previously unrecognized genes that hold substantial promise as future therapeutic targets. This ambitious research endeavor received vital financial backing from prominent federal agencies, including the National Institute on Aging and the National Cancer Institute, underscoring the national priority placed on unraveling complex neurodegenerative conditions.
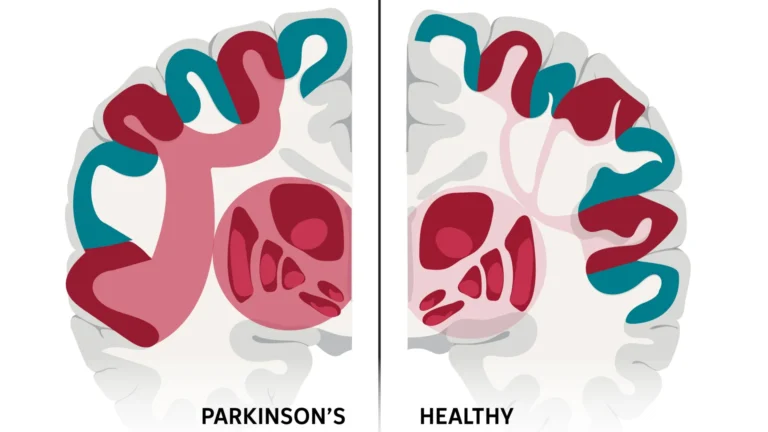
The profound significance of comprehending gene regulatory networks in the context of Alzheimer’s disease cannot be overstated. As the predominant cause of dementia, Alzheimer’s disease presents a formidable and escalating public health challenge. Projections indicate that by the year 2060, nearly 14 million individuals in the United States will be grappling with this debilitating condition. While a growing number of genetic risk factors have been identified, such as variations in the APOE and APP genes, the precise mechanisms through which these genetic predispositions disrupt normal brain function remain largely enigmatic.
Dr. Min Zhang, a co-corresponding author of the study and a distinguished professor of epidemiology and biostatistics, emphasized the limitations of previous research. "The distinct roles played by various brain cell types in the progression of Alzheimer’s disease have been acknowledged, yet the intricate molecular dialogues between them have remained elusive," Dr. Zhang explained. "Our research endeavors to rectify this knowledge gap by furnishing cell-type-specific blueprints of gene regulation within the Alzheimer’s brain. This marks a pivotal shift, moving the field beyond merely observing correlations to actively uncovering the causal mechanisms that propel disease advancement."
The construction of these highly granular gene control maps was facilitated by the analysis of single-cell molecular data. This invaluable dataset was meticulously compiled from brain tissue samples generously donated by 272 participants enrolled in longitudinal aging studies, specifically the Religious Orders Study and the Rush Memory and Aging Project. SIGNET, the computational engine driving this discovery, was conceived as a robust and high-performance computing system. Its architecture is designed to synergistically integrate data derived from single-cell RNA sequencing, which captures the gene expression profiles of individual cells, with whole-genome sequencing data, providing a comprehensive genetic blueprint. This dual-data integration strategy empowers SIGNET to accurately identify cause-and-effect relationships among genes across the entirety of the genome.
Through this sophisticated analytical approach, the researchers were able to reconstruct causal gene regulatory networks for six principal categories of brain cells. This capability allowed them to delineate which genes are most likely acting as molecular conductors, dictating the activity levels of other genes – a feat that conventional correlation-based analytical methods struggle to achieve with reliable accuracy.
Dr. Dabao Zhang, also a co-corresponding author and professor of epidemiology and biostatistics, elaborated on the limitations of existing tools. "The majority of gene-mapping platforms can illustrate which genes exhibit synchronized expression, but they are incapable of definitively determining which genes are truly instigating these changes," Dr. Zhang stated. "Furthermore, certain methodologies impose restrictive assumptions, such as disregarding the complex feedback loops that frequently occur between genes. Our innovative approach leverages the intrinsic information encoded within the DNA sequence itself, thereby enabling the precise identification of authentic cause-and-effect relationships among genes within the brain."
A particularly striking revelation from the study pertains to excitatory neurons, the primary neuronal subtype responsible for transmitting activating signals within the brain. In these critical cells, the researchers uncovered evidence of extensive genetic "rewiring" as Alzheimer’s disease progresses, with nearly 6,000 identified cause-and-effect interactions pointing to significant molecular perturbations.
The investigation also pinpointed hundreds of "hub genes," which function as central regulatory nodes within these networks. These hub genes exert a substantial influence over a multitude of other genes, suggesting their pivotal role in the cascade of detrimental changes observed in the Alzheimer’s brain. The identification of these hub genes opens promising avenues for the development of earlier diagnostic markers and novel therapeutic interventions. Moreover, the study shed new light on the regulatory functions of well-established genes implicated in Alzheimer’s, such as APP. The research demonstrated that APP exerts a potent influence over the expression of other genes, particularly within inhibitory neurons, revealing a previously uncharacterized facet of its molecular activity in the disease.
To ensure the robustness and validity of their findings, the research team subjected their conclusions to rigorous validation. This process involved analyzing an independent cohort of human brain samples, separate from the initial dataset. This independent confirmation significantly bolsters confidence in the observed gene relationships, attesting to their authenticity as genuine biological mechanisms operative in Alzheimer’s disease pathology.
The implications of the SIGNET platform extend far beyond the immediate scope of Alzheimer’s disease research. The sophisticated analytical capabilities of SIGNET hold immense potential for advancing our understanding of a wide spectrum of complex diseases. Researchers anticipate that this framework can be effectively applied to unravel the genetic intricacies of other devastating conditions, including various forms of cancer, autoimmune disorders that compromise the immune system, and a range of mental health conditions that profoundly impact cognitive and emotional well-being.