A groundbreaking scientific revelation has illuminated a critical vulnerability within bacteria, detailing how viruses that prey on these microorganisms have evolved a sophisticated mechanism to dismantle the bacterial cell wall. This intricate molecular sabotage targets MurJ, a protein indispensable for the structural integrity of bacterial envelopes. Astonishingly, the research indicates that disparate viral lineages have independently developed distinct proteins, all converging on the same precise method to incapacitate MurJ. This shared strategy against a vital bacterial component presents a compelling and novel avenue for the development of next-generation antibiotics, offering a glimmer of hope in the escalating global battle against drug-resistant pathogens.
The study, meticulously detailed in the esteemed scientific journal Nature on February 26th, was spearheaded by Yancheng Evelyn Li, a dedicated graduate student operating within the esteemed laboratory of Professor Bil Clemons at the California Institute of Technology (Caltech). Professor Clemons, who holds the distinguished Arthur and Marian Hanisch Memorial Professorship in Biochemistry, served as the corresponding author for this pivotal research.
The escalating threat posed by antibiotic-resistant bacteria, commonly referred to as "superbugs," has reached a critical juncture, necessitating an urgent and innovative approach to therapeutic development. Bacteria, by their very nature, possess an extraordinary capacity for rapid evolution, a characteristic that has unfortunately translated into an alarming acceleration of resistance to existing antimicrobial agents. As Professor Clemons articulated, "Evolution is a formidable force, and within the bacterial realm, the development of resistance to antibiotics transpires with remarkable swiftness. Consequently, we now confront bacterial strains that exhibit imperviousness to the entirety of our current medicinal arsenal." The stark reality of this crisis is underscored by the sobering statistics in the United States alone, where tens of thousands of lives are lost annually to infections caused by antibiotic-resistant bacteria, a figure that is alarmingly on the rise. This grim panorama underscores the profound and immediate need for novel antimicrobial agents capable of effectively combating these resilient pathogens. In response to the diminishing efficacy of conventional drugs, the scientific community is actively engaged in a rigorous search for entirely new points of weakness within bacterial physiology.
A long-standing area of intense scientific scrutiny has been the intricate biochemical pathway that bacteria employ for the construction of peptidoglycan, a rigid polymer that forms the essential structural framework of their cell walls. This critical process, known as the peptidoglycan biosynthesis pathway, represents a particularly attractive target for antimicrobial intervention. The inherent advantage of targeting this pathway lies in the fact that peptidoglycan is a molecular edifice unique to bacterial cells, absent entirely from the cellular composition of human and other eukaryotic organisms. Professor Clemons emphasized this key distinction, stating, "Peptidoglycan stands as a unique characteristic of bacteria, rendering it a highly desirable target for antibiotic development." Indeed, several established antibiotics already exert their therapeutic effects by interfering with this vital pathway. Penicillin, the serendipitous discovery of Alexander Fleming in the mid-20th century, functions by inhibiting a late-stage enzymatic step in peptidoglycan production, a mechanism emulated by related drugs such as amoxicillin.
Central to the successful transport of peptidoglycan precursors across the bacterial inner membrane are three essential protein machinery components: MraY, MurG, and MurJ. These proteins play a crucial role in facilitating the movement of the molecular building blocks required for cell wall assembly from the intracellular environment to the exterior, where the cell wall is constructed. The failure of any single one of these proteins to perform its function would result in the cessation of peptidoglycan synthesis, ultimately leading to bacterial cell death. This fundamental dependency positions them as highly promising targets for therapeutic exploitation. While a substantial body of knowledge exists regarding the functional roles of these proteins, Professor Clemons noted that crucial mechanistic details still remain elusive, representing gaps in our comprehensive understanding.
Currently, no approved pharmaceutical agents directly inhibit the activity of these three pivotal proteins. Nevertheless, Professor Clemons expressed optimism regarding their therapeutic potential, observing, "We are aware that small molecules, whether derived from natural sources or synthesized within chemical libraries, possess the capability to inhibit these proteins. Excitingly, recent discoveries have revealed that bacteriophages, viruses that infect bacteria, have already devised strategies to target this very pathway."
Bacteriophages, often abbreviated as phages, are a diverse group of viruses that specifically infect bacterial cells. For these viral entities to complete their life cycle, they must successfully penetrate a host bacterium, replicate their genetic material within the cell, and subsequently egress to infect other bacterial populations. The critical step of exiting the bacterial cell necessitates traversing the formidable peptidoglycan layer. Professor Clemons elaborated on this necessity, explaining, "The process of re-emergence from the host bacterium requires the phage to overcome the peptidoglycan layer. Given that this layer functions much like a suit of armor, phages can become ensnared if they are unable to breach it." The research conducted by Professor Clemons’ lab specifically focuses on small bacteriophages characterized by their single-stranded DNA or RNA genomes. These viruses, with their compact genetic material, rely on elegantly simple strategies to achieve bacterial lysis. In a significant prior publication in the journal Science in 2023, the team reported on their investigations into the well-studied phage φX174, a virus with a rich history of research at Caltech.
Small bacteriophages employ specialized protein effectors, known as single-gene lysis proteins, or Sgls, to induce bacterial cell death. The work undertaken by Li and Clemons has centered on Sgls that specifically target MurJ, one of the aforementioned key proteins involved in cell wall construction. MurJ functions as a flippase, a molecular transporter that facilitates the movement of peptidoglycan precursor molecules from the inner leaflet of the bacterial membrane to the outer leaflet, thereby making them accessible for incorporation into the growing cell wall. Previous research conducted by collaborators had already established that two distinct and evolutionarily unrelated Sgls, SglM and SglPP7, both achieve bacterial inactivation by effectively blocking the function of MurJ.
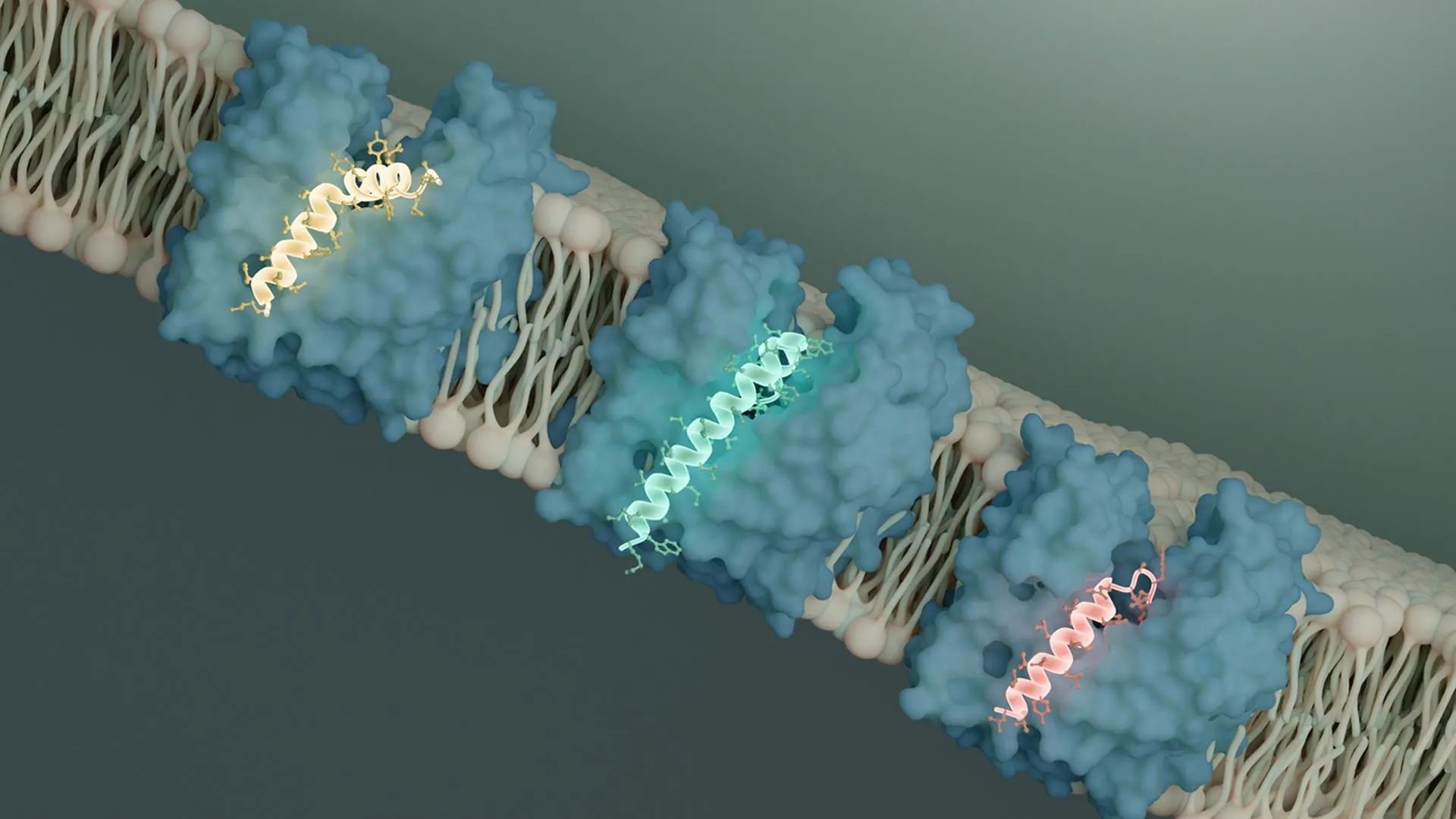
To elucidate the precise molecular mechanisms underlying this viral intervention, Li utilized state-of-the-art cryo-electron microscopy (cryo-EM) techniques at Caltech’s Beckman Institute Biological and Cryogenic Transmission Electron Microscopy (Cryo-EM) Resource Center. Flippases, such as MurJ, operate by undergoing conformational changes to shuttle molecules across the membrane without creating a permanent pore. When MurJ binds to its substrate within the cell, it adopts a specific three-dimensional shape that facilitates the release of the cargo on the exterior side of the membrane. Through her meticulous cryo-EM analysis, Li determined that both SglM and SglPP7 bind to a specific groove on the MurJ protein, effectively preventing the conformational shift essential for its transport function.
"It is unequivocally clear that both of these Sgls engage with MurJ in an outward-facing conformation, effectively locking it into this particular orientation," stated Li. This observation holds significant promise for therapeutic development because the outward-facing conformation of MurJ is more exposed to the external environment of the bacterial cell, potentially rendering it a more accessible target for future drug molecules compared to an inward-facing conformation.
The discovery that two unrelated viral proteins have independently evolved to inhibit MurJ in such a similar fashion was a remarkable and unexpected finding. Professor Clemons commented on the significance of this observation, noting, "These peptides, which share no discernible evolutionary lineage, have both independently devised strategies to target MurJ through remarkably similar mechanisms. This represents a striking example of convergent evolution, where distinct evolutionary trajectories converge upon the same functional solution. We were genuinely taken by surprise!"
Given the accelerated pace of viral evolution, the research team posits that a vast number of other bacteriophages likely harbor similar Sgl proteins. The relative ease with which bacteriophages can be isolated and their genomes sequenced offers a rich avenue for uncovering additional biological insights and identifying novel antimicrobial targets. In their Nature publication, the researchers, in collaboration with a colleague, further investigated this phenomenon by analyzing the genome of yet another phage. This analysis led to the identification of a novel Sgl, designated SglCJ3 (derived from a phage genome predicted to be named Changjiang3), which was subsequently subjected to structural analysis using cryo-EM. Li successfully determined the structure of SglCJ3 in complex with MurJ, confirming that this third viral protein also functions by locking MurJ into the same outward-facing conformation.
"This represents a third independent evolutionary event where a distinct peptide has evolved to inhibit the same target through a comparable mechanism," Professor Clemons remarked. "This provides the first compelling evidence that nature itself recognizes MurJ as an exceptional target for bacterial inactivation, strongly suggesting that we should follow this evolutionary lead and pursue the development of therapeutics that target MurJ. This research beautifully illustrates the profound capacity of fundamental biological inquiry to contribute solutions to pressing medical challenges. Our future endeavors are now firmly directed towards leveraging the discoveries of Sgl proteins, and we are hopeful for continued support to translate these foundational concepts into tangible clinical realities."
The foundational scientific paper detailing these findings is titled "Convergent MurJ flippase inhibition by phage lysis proteins." The authorship of this significant work includes Professor Bil Clemons and Yancheng Evelyn Li of Caltech, alongside Caltech graduate student Grace F. Baron. The research was further enriched by the contributions of Francesca S. Antillon, Karthik Chamakura, and Ry Young from Texas A&M University. Financial support for this groundbreaking research was generously provided by the Chan Zuckerberg Initiative, the National Institutes of Health, the G. Harold and Leila Y. Mathers Foundation, and the Center for Phage Technology at Texas A&M, an initiative jointly sponsored by Texas A&M AgriLife.