An international consortium of scientists, spearheaded by researchers at Durham University and involving collaborators from Iceland, Norway, and Poland, has unveiled a suite of previously unrecognized DNA-binding proteins. These remarkable molecules, isolated from some of the planet’s most inhospitable environments, including the superheated waters of Icelandic volcanic lakes and the crushing pressures of deep-sea hydrothermal vents located over two kilometers beneath the surface of the North Atlantic Ocean, have demonstrated a significant capacity to enhance the speed and accuracy of rapid medical diagnostic assays for infectious diseases.
The exploration of Earth’s extreme environments represents a frontier in the quest for novel biological tools, offering a vast, largely untapped reservoir of unique enzymes and proteins. Nature, in its evolutionary resilience, has cultivated organisms capable of thriving under conditions that would be lethal to most life forms, and these organisms often possess biochemical machinery adapted to such challenges. To systematically probe this genetic diversity, the research team employed advanced next-generation DNA sequencing technologies. This approach allowed them to sift through enormous digital archives, encompassing millions of potential protein sequences, effectively mining nature’s genetic library for hidden treasures.
Through meticulous analysis of this extensive genetic data, the scientists successfully identified a class of novel proteins exhibiting a pronounced affinity for single-stranded DNA. What set these proteins apart was their extraordinary inherent stability, allowing them to function effectively and retain their structural integrity under a formidable array of environmental stressors. These included exceptionally high temperatures, extreme fluctuations in pH levels, and environments with exceptionally high salt concentrations – conditions that typically denature or inactivate most common proteins. This resilience is a key indicator of their potential utility in demanding biotechnological applications.
Further detailed investigation into these newly discovered proteins involved a comprehensive suite of sophisticated laboratory techniques. These studies confirmed their exceptional robustness, particularly their remarkable thermal stability. This characteristic is a critical attribute for proteins intended for use in biotechnology and advanced medical diagnostics, where reactions often occur at elevated temperatures or require components that can withstand harsh processing conditions. The ability of these proteins to maintain their functional shape and activity under such duress makes them highly attractive candidates for further development.
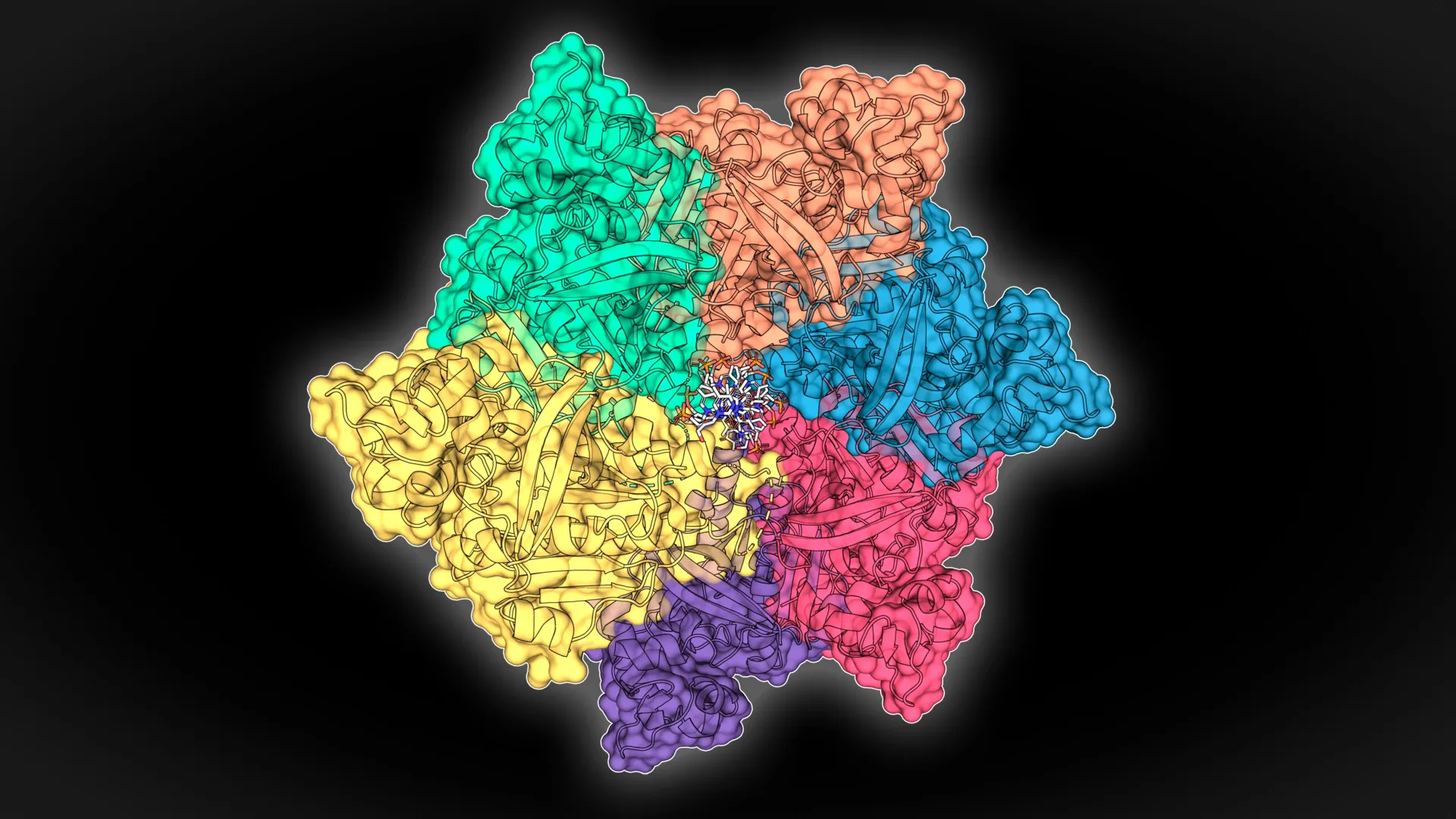
Beyond their functional resilience, researchers also succeeded in elucidating the three-dimensional atomic structures of these proteins with high resolution. This detailed structural information is invaluable, as it provides profound insights into the molecular mechanisms underlying their function. Understanding precisely how these proteins fold and interact with their targets, such as DNA, not only explains their observed stability but also paves the way for rational protein engineering. Through targeted modifications guided by structural data, scientists can potentially refine these proteins further, optimizing their properties for specific applications or even creating entirely new functionalities.
A particularly exciting demonstration of the practical utility of these extremophilic proteins came with their incorporation into rapid diagnostic tests that utilize loop-mediated isothermal amplification, commonly known as LAMP. LAMP assays are a powerful class of molecular diagnostic tools that can detect the genetic material of viruses, bacteria, or parasites. Crucially, they achieve this without the need for sophisticated laboratory equipment such as thermal cyclers, making them ideal for point-of-care testing, resource-limited settings, or rapid field diagnostics. These tests operate at a constant temperature, simplifying their implementation.
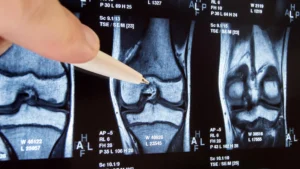
When one of the newly identified DNA-binding proteins was introduced into the LAMP assay, a significant enhancement in performance was observed. The diagnostic tests not only became considerably faster, reducing the time required to obtain a result, but also exhibited increased sensitivity. This heightened sensitivity translates directly into a greater ability to detect minute quantities of pathogen genetic material. The researchers demonstrated this improvement by successfully detecting viral RNA from pathogens like SARS-CoV-2, the virus responsible for COVID-19, as well as DNA from a range of other infectious agents, with greater reliability. These findings underscore the profound potential of exploring Earth’s most extreme habitats as a source for developing innovative biological tools that can address pressing global health challenges.
Professor Ehmke Pohl, the lead investigator for the study from Durham University, emphasized the broader implications of this work. "This research powerfully illustrates the immense potential residing in bioprospecting endeavors focused on extreme environments," he stated. "The outcomes are not merely significant for the burgeoning bioeconomy; they also lay a foundational contribution to all artificial intelligence (AI) methods that are increasingly being employed in the critical fields of protein structure prediction and protein design." This connection to AI highlights how real-world biological data from diverse sources can fuel advancements in computational biology.
The biotechnology sector is in perpetual pursuit of enzymes and proteins that can perform reliably and efficiently under demanding operational conditions. Proteins that naturally originate from environments like geothermal hot springs or deep-sea hydrothermal vents are especially prized because they have evolved mechanisms to function optimally within these harsh settings. This inherent robustness offers a significant advantage over proteins engineered from less resilient organisms, potentially reducing the need for extensive and costly stabilization efforts.
Furthermore, these discoveries hold promise for advancing the field of protein prediction and design, particularly in conjunction with artificial intelligence. AI systems designed to model and predict protein structures and functions thrive on vast, diverse, and high-quality datasets of real biological examples. The inclusion of data from these newly characterized extremophilic proteins enriches these datasets, providing AI models with novel structural motifs and functional insights that can improve their predictive accuracy and generative capabilities in designing new proteins with desired characteristics.
The research team is not resting on its laurels; their investigation into additional DNA-binding proteins from extreme environments is ongoing. Several highly promising candidates are already under scrutiny, suggesting that this discovery may be just the tip of the iceberg. Concurrently, efforts are underway to develop improved versions of the already identified proteins through protein engineering. The team is also actively designing new LAMP assays specifically tailored to detect pathogens responsible for neglected tropical diseases, including leishmaniasis and Chagas disease, thereby aiming to expand the reach of accessible diagnostics to underserved populations. These developmental efforts are being conducted in close collaboration with colleagues within Durham University’s Department of Biosciences.
To explore the commercial viability of these groundbreaking discoveries, the research group has partnered with ArcticZymes, a Norwegian biotechnology company. This collaboration is focused on identifying and developing potential market applications for the newly identified proteins, aiming to translate scientific breakthroughs into tangible solutions for industry and healthcare. The synergy between academic research and commercial enterprise is crucial for bringing such innovative technologies from the laboratory to the global stage.